-
 The Therapeutic Mechanisms of Mesenchymal Stem Cells in MS—A Review Focusing on Neuroprotective Properties
The Therapeutic Mechanisms of Mesenchymal Stem Cells in MS—A Review Focusing on Neuroprotective Properties -
 The Bee Gut Microbiota: Bridging Infective Agents Potential in the One Health Context
The Bee Gut Microbiota: Bridging Infective Agents Potential in the One Health Context -
 The Role of Extracellular Vesicles in Atopic Dermatitis
The Role of Extracellular Vesicles in Atopic Dermatitis -
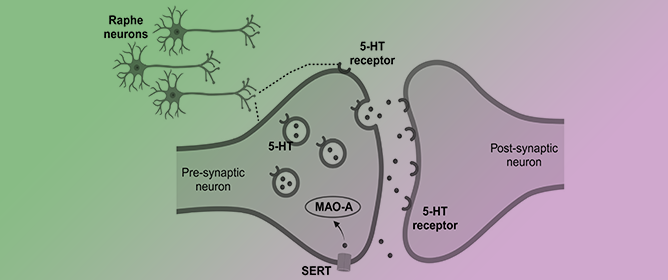 Revisiting the Role of Serotonin in Sleep-Disordered Breathing
Revisiting the Role of Serotonin in Sleep-Disordered Breathing -
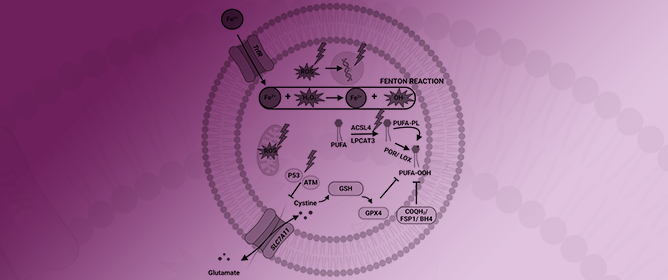 Ferroptosis: Frenemy of Radiotherapy
Ferroptosis: Frenemy of Radiotherapy
Journal Description
International Journal of Molecular Sciences
International Journal of Molecular Sciences
is an international, peer-reviewed, open access journal providing an advanced forum for biochemistry, molecular and cell biology, molecular biophysics, molecular medicine, and all aspects of molecular research in chemistry, and is published semimonthly online by MDPI. The Australian Society of Plant Scientists (ASPS), Epigenetics Society, European Calcium Society (ECS), European Chitin Society (EUCHIS), Spanish Society for Cell Biology (SEBC) and others are affiliated with IJMS and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, MEDLINE, Embase, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q1 (Biochemistry & Molecular Biology) / CiteScore - Q1 (Inorganic Chemistry)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.3 days after submission; acceptance to publication is undertaken in 2.6 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about the IJMS.
- Companion journals for IJMS include: Biophysica, Obesities, Stresses and Lymphatics.
Impact Factor:
5.6 (2022);
5-Year Impact Factor:
6.2 (2022)
Latest Articles
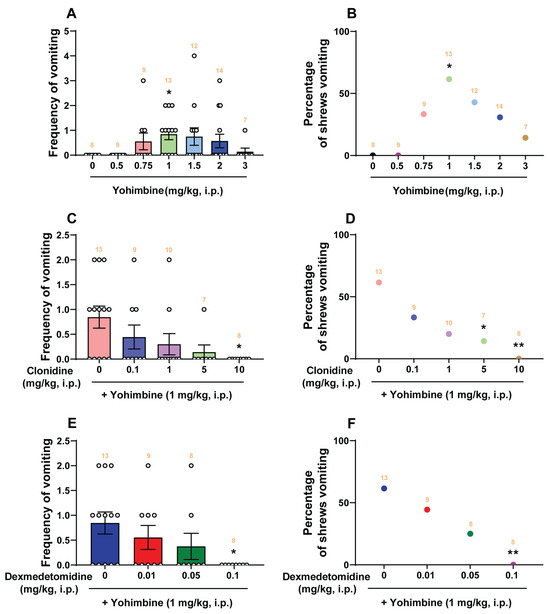
A Comparative Study of the Antiemetic Effects of α2-Adrenergic Receptor Agonists Clonidine and Dexmedetomidine against Diverse Emetogens in the Least Shrew (Cryptotis parva) Model of Emesis
Int. J. Mol. Sci. 2024, 25(9), 4603; https://doi.org/10.3390/ijms25094603 (registering DOI) - 23 Apr 2024
Abstract
In contrast to cats and dogs, here we report that the α2-adrenergic receptor antagonist yohimbine is emetic and corresponding agonists clonidine and dexmedetomidine behave as antiemetics in the least shrew model of vomiting. Yohimbine (0, 0.5, 0.75, 1, 1.5, 2, and
[...] Read more.
In contrast to cats and dogs, here we report that the α2-adrenergic receptor antagonist yohimbine is emetic and corresponding agonists clonidine and dexmedetomidine behave as antiemetics in the least shrew model of vomiting. Yohimbine (0, 0.5, 0.75, 1, 1.5, 2, and 3 mg/kg, i.p.) caused vomiting in shrews in a bell-shaped and dose-dependent manner, with a maximum frequency (0.85 ± 0.22) at 1 mg/kg, which was accompanied by a key central contribution as indicated by increased expression of c-fos, serotonin and substance P release in the shrew brainstem emetic nuclei. Our comparative study in shrews demonstrates that clonidine (0, 0.1, 1, 5, and 10 mg/kg, i.p.) and dexmedetomidine (0, 0.01, 0.05, and 0.1 mg/kg, i.p.) not only suppress yohimbine (1 mg/kg, i.p.)-evoked vomiting in a dose-dependent manner, but also display broad-spectrum antiemetic effects against diverse well-known emetogens, including 2-Methyl-5-HT, GR73632, McN-A-343, quinpirole, FPL64176, SR141716A, thapsigargin, rolipram, and ZD7288. The antiemetic inhibitory ID50 values of dexmedetomidine against the evoked emetogens are much lower than those of clonidine. At its antiemetic doses, clonidine decreased shrews’ locomotor activity parameters (distance moved and rearing), whereas dexmedetomidine did not do so. The results suggest that dexmedetomidine represents a better candidate for antiemetic potential with advantages over clonidine.
Full article
(This article belongs to the Special Issue Intercellular and Intracellular Communication in Human Health and Disease 2.0)
►
Show Figures
Open AccessArticle
Modulating Endoplasmic Reticulum Chaperones and Mutant Protein Degradation in GABRG2(Q390X) Associated with Genetic Epilepsy with Febrile Seizures Plus and Dravet Syndrome
by
Sarah Poliquin, Gerald Nwosu, Karishma Randhave, Wangzhen Shen, Carson Flamm and Jing-Qiong Kang
Int. J. Mol. Sci. 2024, 25(9), 4601; https://doi.org/10.3390/ijms25094601 (registering DOI) - 23 Apr 2024
Abstract
A significant number of patients with genetic epilepsy do not obtain seizure freedom, despite developments in new antiseizure drugs, suggesting a need for novel therapeutic approaches. Many genetic epilepsies are associated with misfolded mutant proteins, including GABRG2(Q390X)-associated Dravet syndrome, which we have
[...] Read more.
A significant number of patients with genetic epilepsy do not obtain seizure freedom, despite developments in new antiseizure drugs, suggesting a need for novel therapeutic approaches. Many genetic epilepsies are associated with misfolded mutant proteins, including GABRG2(Q390X)-associated Dravet syndrome, which we have previously shown to result in intracellular accumulation of mutant GABAA receptor γ2(Q390X) subunit protein. Thus, a potentially promising therapeutic approach is modulation of proteostasis, such as increasing endoplasmic reticulum (ER)-associated degradation (ERAD). To that end, we have here identified an ERAD-associated E3 ubiquitin ligase, HRD1, among other ubiquitin ligases, as a strong modulator of wildtype and mutant γ2 subunit expression. Overexpressing HRD1 or knockdown of HRD1 dose-dependently reduced the γ2(Q390X) subunit. Additionally, we show that zonisamide (ZNS)—an antiseizure drug reported to upregulate HRD1—reduces seizures in the Gabrg2+/Q390X mouse. We propose that a possible mechanism for this effect is a partial rescue of surface trafficking of GABAA receptors, which are otherwise sequestered in the ER due to the dominant-negative effect of the γ2(Q390X) subunit. Furthermore, this partial rescue was not due to changes in ER chaperones BiP and calnexin, as total expression of these chaperones was unchanged in γ2(Q390X) models. Our results here suggest that leveraging the endogenous ERAD pathway may present a potential method to degrade neurotoxic mutant proteins like the γ2(Q390X) subunit. We also demonstrate a pharmacological means of regulating proteostasis, as ZNS alters protein trafficking, providing further support for the use of proteostasis regulators for the treatment of genetic epilepsies.
Full article
(This article belongs to the Special Issue Epilepsy: From Molecular Basis to Therapy)
►▼
Show Figures
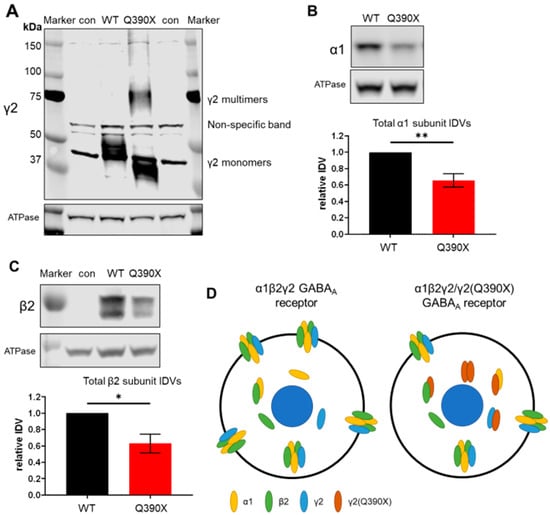
Figure 1
Open AccessArticle
HOPS/TMUB1 Enhances Apoptosis in TP53 Mutation-Independent Setting in Human Cancers
by
Nicola Di-Iacovo, Simona Ferracchiato, Stefania Pieroni, Damiano Scopetti, Marilena Castelli, Danilo Piobbico, Luca Pierucci, Marco Gargaro, Davide Chiasserini, Giuseppe Servillo and Maria Agnese Della-Fazia
Int. J. Mol. Sci. 2024, 25(9), 4600; https://doi.org/10.3390/ijms25094600 (registering DOI) - 23 Apr 2024
Abstract
TP53 mutations are prevalent in various cancers, yet the complexity of apoptotic pathway deregulation suggests the involvement of additional factors. HOPS/TMUB1 is known to extend the half-life of p53 under normal and stress conditions, implying a regulatory function. This study investigates, for the
[...] Read more.
TP53 mutations are prevalent in various cancers, yet the complexity of apoptotic pathway deregulation suggests the involvement of additional factors. HOPS/TMUB1 is known to extend the half-life of p53 under normal and stress conditions, implying a regulatory function. This study investigates, for the first time, the potential modulatory role of the ubiquitin-like-protein HOPS/TMUB1 in p53-mutants. A comprehensive analysis of apoptosis in the most frequent p53-mutants, R175, R248, and R273, in SKBR3, MIA PaCa2, and H1975 cells indicates that the overexpression of HOPS induces apoptosis at least equivalent to that caused by DNA damage. Immunoprecipitation assays confirm HOPS binding to p53-mutant forms. The interaction of HOPS/TMUB1 with p53-mutants strengthens its effect on the apoptotic cascade, showing a context-dependent gain or loss of function. Gene expression analysis of the MYC and TP63 genes shows that H1975 exhibit a gain-of-function profile, while SKBR3 promote apoptosis in a TP63-dependent manner. The TCGA data further corroborate HOPS/TMUB1’s positive correlation with apoptotic genes BAX, BBC3, and NOXA1, underscoring its relevance in patient samples. Notably, singular TP53 mutations inadequately explain pathway dysregulation, emphasizing the need to explore additional contributing factors. These findings illuminate the intricate interplay among TP53 mutations, HOPS/TMUB1, and apoptotic pathways, providing valuable insights for targeted cancer interventions.
Full article
(This article belongs to the Special Issue Novel Therapeutic Targets in Cancers 2.0)
►▼
Show Figures
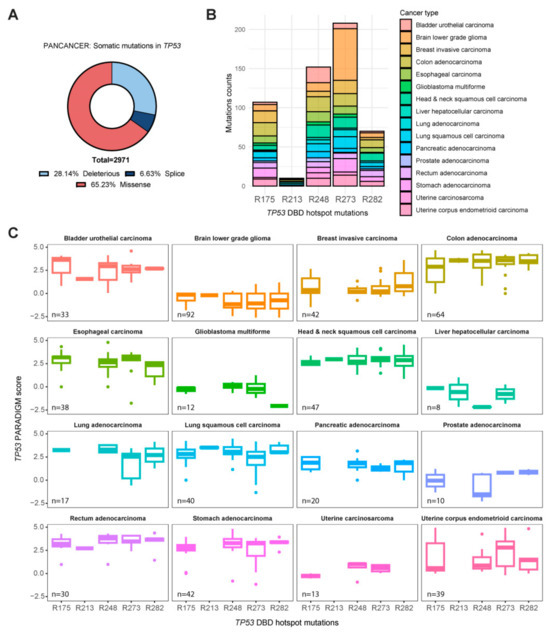
Figure 1
Open AccessReview
The Impact of the Aryl Hydrocarbon Receptor on Antenatal Chemical Exposure-Induced Cardiovascular–Kidney–Metabolic Programming
by
You-Lin Tain and Chien-Ning Hsu
Int. J. Mol. Sci. 2024, 25(9), 4599; https://doi.org/10.3390/ijms25094599 (registering DOI) - 23 Apr 2024
Abstract
Early life exposure lays the groundwork for the risk of developing cardiovascular–kidney–metabolic (CKM) syndrome in adulthood. Various environmental chemicals to which pregnant mothers are commonly exposed can disrupt fetal programming, leading to a wide range of CKM phenotypes. The aryl hydrocarbon receptor (AHR)
[...] Read more.
Early life exposure lays the groundwork for the risk of developing cardiovascular–kidney–metabolic (CKM) syndrome in adulthood. Various environmental chemicals to which pregnant mothers are commonly exposed can disrupt fetal programming, leading to a wide range of CKM phenotypes. The aryl hydrocarbon receptor (AHR) has a key role as a ligand-activated transcription factor in sensing these environmental chemicals. Activating AHR through exposure to environmental chemicals has been documented for its adverse impacts on cardiovascular diseases, hypertension, diabetes, obesity, kidney disease, and non-alcoholic fatty liver disease, as evidenced by both epidemiological and animal studies. In this review, we compile current human evidence and findings from animal models that support the connection between antenatal chemical exposures and CKM programming, focusing particularly on AHR signaling. Additionally, we explore potential AHR modulators aimed at preventing CKM syndrome. As the pioneering review to present evidence advocating for the avoidance of toxic chemical exposure during pregnancy and deepening our understanding of AHR signaling, this has the potential to mitigate the global burden of CKM syndrome in the future.
Full article
(This article belongs to the Special Issue Molecular Insights into the Developmental Origins of Health and Disease)
►▼
Show Figures
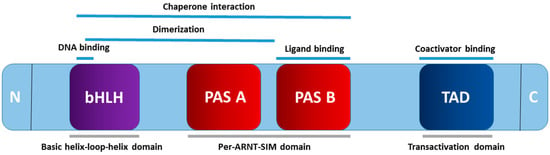
Figure 1
Open AccessArticle
Genetic Landscape and Clinical Features of Hyperphenylalaninemia in North Ossetia-Alania: High Frequency of P281L and P211T Genetic Variants in the PAH Gene
by
Inna S. Tebieva, Polina V. Mishakova, Yulia V. Gabisova, Alana V. Khokhova, Tamara G. Kaloeva, Andrey V. Marakhonov, Olga A. Shchagina, Alexander V. Polyakov, Evgeny K. Ginter, Sergey I. Kutsev and Rena A. Zinchenko
Int. J. Mol. Sci. 2024, 25(9), 4598; https://doi.org/10.3390/ijms25094598 (registering DOI) - 23 Apr 2024
Abstract
This study, conducted in the Republic of North Ossetia-Alania (RNOA), aimed to explore the genetic landscape of hyperphenylalaninemia (HPA) and phenylketonuria (PKU) in the Ossetian population using data from newborn screening (NBS). Through comprehensive molecular genetic analysis of 29 patients with HPA from
[...] Read more.
This study, conducted in the Republic of North Ossetia-Alania (RNOA), aimed to explore the genetic landscape of hyperphenylalaninemia (HPA) and phenylketonuria (PKU) in the Ossetian population using data from newborn screening (NBS). Through comprehensive molecular genetic analysis of 29 patients with HPA from diverse ethnic backgrounds, two major genetic variants in the PAH gene, P281L and P211T, were identified, constituting 50% of all detected pathogenic alleles in Ossetian patients. Remarkably, these variants exhibited an exceptionally high frequency in the Ossetian population, surpassing global prevalence rates. This study unveiled a notable prevalence of mild forms of HPA (78%), underscoring the importance of genetic counseling for carriers of pathogenic variants in the PAH gene. Moreover, the findings emphasized the necessity for ongoing monitoring of patients with mild forms, as they may lack significant symptoms for diagnosis, potentially impacting offspring. Overall, this research offers valuable insights into the genetic landscape of HPA and PKU in the Ossetian population.
Full article
(This article belongs to the Special Issue Advances in Human Hereditary Diseases: Genetics and Genomics Research)
Open AccessArticle
Microcephaly Gene Mcph1 Deficiency Induces p19ARF-Dependent Cell Cycle Arrest and Senescence
by
Yi-Nan Jiang, Yizhen Gao, Xianxin Lai, Xinjie Li, Gen Liu, Mingmei Ding, Zhiyi Wang, Zixiang Guo, Yinying Qin, Xin Li, Litao Sun, Zhao-Qi Wang and Zhong-Wei Zhou
Int. J. Mol. Sci. 2024, 25(9), 4597; https://doi.org/10.3390/ijms25094597 (registering DOI) - 23 Apr 2024
Abstract
MCPH1 has been identified as the causal gene for primary microcephaly type 1, a neurodevelopmental disorder characterized by reduced brain size and delayed growth. As a multifunction protein, MCPH1 has been reported to repress the expression of TERT and interact with transcriptional regulator
[...] Read more.
MCPH1 has been identified as the causal gene for primary microcephaly type 1, a neurodevelopmental disorder characterized by reduced brain size and delayed growth. As a multifunction protein, MCPH1 has been reported to repress the expression of TERT and interact with transcriptional regulator E2F1. However, it remains unclear whether MCPH1 regulates brain development through its transcriptional regulation function. This study showed that the knockout of Mcph1 in mice leads to delayed growth as early as the embryo stage E11.5. Transcriptome analysis (RNA-seq) revealed that the deletion of Mcph1 resulted in changes in the expression levels of a limited number of genes. Although the expression of some of E2F1 targets, such as Satb2 and Cdkn1c, was affected, the differentially expressed genes (DEGs) were not significantly enriched as E2F1 target genes. Further investigations showed that primary and immortalized Mcph1 knockout mouse embryonic fibroblasts (MEFs) exhibited cell cycle arrest and cellular senescence phenotype. Interestingly, the upregulation of p19ARF was detected in Mcph1 knockout MEFs, and silencing p19Arf restored the cell cycle and growth arrest to wild-type levels. Our findings suggested it is unlikely that MCPH1 regulates neurodevelopment through E2F1-mediated transcriptional regulation, and p19ARF-dependent cell cycle arrest and cellular senescence may contribute to the developmental abnormalities observed in primary microcephaly.
Full article
(This article belongs to the Special Issue Molecular Research in Neurodevelopmental Disorders)
►▼
Show Figures
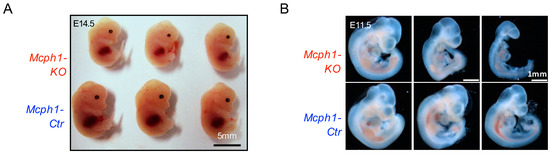
Figure 1
Open AccessArticle
RNA-Seq Bulked Segregant Analysis of an Exotic B. napus ssp. napobrassica (Rutabaga) F2 Population Reveals Novel QTLs for Breeding Clubroot-Resistant Canola
by
Zhiyu Yu, Rudolph Fredua-Agyeman, Stephen E. Strelkov and Sheau-Fang Hwang
Int. J. Mol. Sci. 2024, 25(9), 4596; https://doi.org/10.3390/ijms25094596 (registering DOI) - 23 Apr 2024
Abstract
In this study, a rutabaga (Brassica napus ssp. napobrassica) donor parent FGRA106, which exhibited broad-spectrum resistance to 17 isolates representing 16 pathotypes of Plasmodiophora brassicae, was used in genetic crosses with the susceptible spring-type canola (B. napus ssp. napus
[...] Read more.
In this study, a rutabaga (Brassica napus ssp. napobrassica) donor parent FGRA106, which exhibited broad-spectrum resistance to 17 isolates representing 16 pathotypes of Plasmodiophora brassicae, was used in genetic crosses with the susceptible spring-type canola (B. napus ssp. napus) accession FG769. The F2 plants derived from a clubroot-resistant F1 plant were screened against three P. brassicae isolates representing pathotypes 3A, 3D, and 3H. Chi-square (χ2) goodness-of-fit tests indicated that the F2 plants inherited two major clubroot resistance genes from the CR donor FGRA106. The total RNA from plants resistant (R) and susceptible (S) to each pathotype were pooled and subjected to bulked segregant RNA-sequencing (BSR-Seq). The analysis of gene expression profiles identified 431, 67, and 98 differentially expressed genes (DEGs) between the R and S bulks. The variant calling method indicated a total of 12 (7 major + 5 minor) QTLs across seven chromosomes. The seven major QTLs included: BnaA5P3A.CRX1.1, BnaC1P3H.CRX1.2, and BnaC7P3A.CRX1.1 on chromosomes A05, C01, and C07, respectively; and BnaA8P3D.CRX1.1, BnaA8P3D.RCr91.2/BnaA8P3H.RCr91.2, BnaA8P3H.Crr11.3/BnaA8P3D.Crr11.3, and BnaA8P3D.qBrCR381.4 on chromosome A08. A total of 16 of the DEGs were located in the major QTL regions, 13 of which were on chromosome C07. The molecular data suggested that clubroot resistance in FGRA106 may be controlled by major and minor genes on both the A and C genomes, which are deployed in different combinations to confer resistance to the different isolates. This study provides valuable germplasm for the breeding of clubroot-resistant B. napus cultivars in Western Canada.
Full article
(This article belongs to the Special Issue Recent Advances in Epigenetics in Plant Research)
►▼
Show Figures
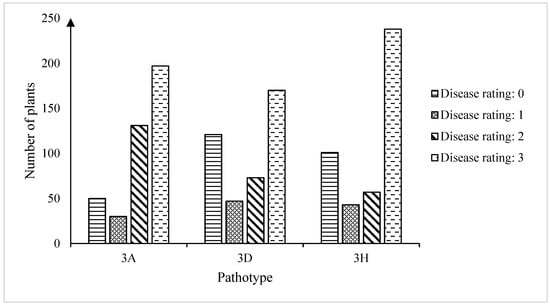
Figure 1
Open AccessArticle
An Effective and Promising Strategy for Plant Protection: Synthesis of L-Carvone-Based Thiazolinone–Hydrazone/Nanochitosan Complexes with Antifungal Activity and Sustained Releasing Performance
by
Baoyu Li, Wengui Duan, Guishan Lin, Xianli Ma, Rongzhu Wen and Zhaolei Zhang
Int. J. Mol. Sci. 2024, 25(9), 4595; https://doi.org/10.3390/ijms25094595 (registering DOI) - 23 Apr 2024
Abstract
The development of novel natural product-derived nano-pesticide systems with loading capacity and sustained releasing performance of bioactive compounds is considered an effective and promising plant protection strategy. In this work, 25 L-carvone-based thiazolinone–hydrazone compounds 4a~4y were synthesized by the multi-step
[...] Read more.
The development of novel natural product-derived nano-pesticide systems with loading capacity and sustained releasing performance of bioactive compounds is considered an effective and promising plant protection strategy. In this work, 25 L-carvone-based thiazolinone–hydrazone compounds 4a~4y were synthesized by the multi-step modification of L-carvone and structurally confirmed. Compound 4h was found to show favorable and broad-spectrum antifungal activity through the in vitro antifungal activity evaluation of compounds 4a~4y against eight phytopathogenic fungi. Thus, it could serve as a leading compound for new antifungal agents in agriculture. Moreover, the L-carvone-based nanochitosan carrier 7 bearing the 1,3,4-thiadiazole-amide group was rationally designed for the loading and sustained releasing applications of compound 4h, synthesized, and characterized. It was proven that carrier 7 had good thermal stability below 200 °C, dispersed well in the aqueous phase to form numerous nanoparticles with a size of~20 nm, and exhibited an unconsolidated and multi-aperture micro-structure. Finally, L-carvone-based thiazolinone–hydrazone/nanochitosan complexes were fabricated and investigated for their sustained releasing behaviors. Among them, complex 7/4h-2 with a well-distributed, compact, and columnar micro-structure displayed the highest encapsulation efficiency and desirable sustained releasing property for compound 4h and thus showed great potential as an antifungal nano-pesticide for further studies.
Full article
(This article belongs to the Section Molecular Plant Sciences)
►▼
Show Figures
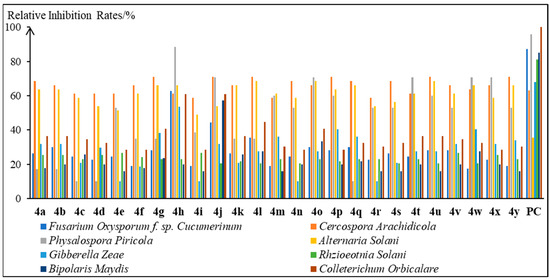
Figure 1
Open AccessArticle
Assessment of the Correlation and Diagnostic Accuracy between Cerebrospinal Fluid and Plasma Alzheimer’s Disease Biomarkers: A Comparison of the Lumipulse and Simoa Platforms
by
Farida Dakterzada, Raffaela Cipriani, Ricard López-Ortega, Alfonso Arias, Iolanda Riba-Llena, Maria Ruiz-Julián, Raquel Huerto, Nuria Tahan, Carlos Matute, Estibaliz Capetillo-Zarate and Gerard Piñol-Ripoll
Int. J. Mol. Sci. 2024, 25(9), 4594; https://doi.org/10.3390/ijms25094594 (registering DOI) - 23 Apr 2024
Abstract
We compared the clinical and analytical performance of Alzheimer’s disease (AD) plasma biomarkers measured using the single-molecule array (Simoa) and Lumipulse platforms. We quantified the plasma levels of amyloid beta 42 (Aβ42), Aβ40, phosphorylated tau (Ptau181), and total tau biomarkers in 81 patients
[...] Read more.
We compared the clinical and analytical performance of Alzheimer’s disease (AD) plasma biomarkers measured using the single-molecule array (Simoa) and Lumipulse platforms. We quantified the plasma levels of amyloid beta 42 (Aβ42), Aβ40, phosphorylated tau (Ptau181), and total tau biomarkers in 81 patients with mild cognitive impairment (MCI), 30 with AD, and 16 with non-AD dementia. We found a strong correlation between the Simoa and Lumipulse methods. Concerning the clinical diagnosis, Simoa Ptau181/Aβ42 (AUC 0.739, 95% CI 0.592–0.887) and Lumipulse Aβ42 and Ptau181/Aβ42 (AUC 0.735, 95% CI 0.589–0.882 and AUC 0.733, 95% CI 0.567–0.900) had the highest discriminating power. However, their power was significantly lower than that of CSF Aβ42/Aβ40, as measured by Lumipulse (AUC 0.879, 95% CI 0.766–0.992). Simoa Ptau181 and Lumipulse Ptau181/Aβ42 were the markers most consistent with the CSF Aβ42/Aβ40 status (AUC 0.801, 95% CI 0.712–0.890 vs. AUC 0.870, 95% CI 0.806–0.934, respectively) at the ≥2.127 and ≥0.084 cut-offs, respectively. The performance of the Simoa and Lumipulse plasma AD assays is weaker than that of CSF AD biomarkers. At present, the analysed AD plasma biomarkers may be useful for screening to reduce the number of lumbar punctures in the clinical setting.
Full article
(This article belongs to the Special Issue New Trends in Alzheimer’s Disease Research: From Molecular Mechanisms to Therapeutics 2.0)
►▼
Show Figures
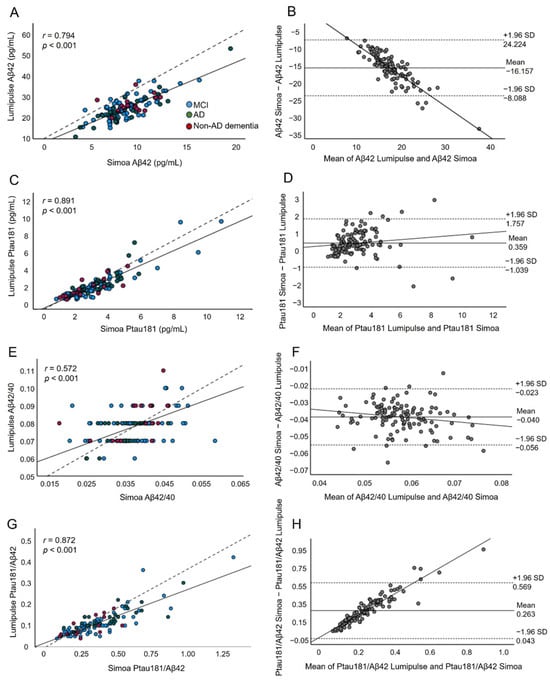
Figure 1
Open AccessArticle
BMP Stimulation Differentially Affects Phosphorylation and Protein Stability of β-Catenin in Breast Cancer Cell Lines
by
Mustafa Ilhan, Nurcan Hastar, Branka Kampfrath, Deniz Neslihan Spierling, Jerome Jatzlau and Petra Knaus
Int. J. Mol. Sci. 2024, 25(9), 4593; https://doi.org/10.3390/ijms25094593 (registering DOI) - 23 Apr 2024
Abstract
Increased expression and nuclear translocation of β-CATENIN is frequently observed in breast cancer, and it correlates with poor prognosis. Current treatment strategies targeting β-CATENIN are not as efficient as desired. Therefore, detailed understanding of β-CATENIN regulation is crucial. Bone morphogenetic proteins (BMP) and
[...] Read more.
Increased expression and nuclear translocation of β-CATENIN is frequently observed in breast cancer, and it correlates with poor prognosis. Current treatment strategies targeting β-CATENIN are not as efficient as desired. Therefore, detailed understanding of β-CATENIN regulation is crucial. Bone morphogenetic proteins (BMP) and Wingless/Integrated (WNT) pathway crosstalk is well-studied for many cancer types including colorectal cancer, whereas it is still poorly understood for breast cancer. Analysis of breast cancer patient data revealed that BMP2 and BMP6 were significantly downregulated in tumors. Since mutation frequency in genes enhancing β-CATENIN protein stability is relatively low in breast cancer, we aimed to investigate whether decreased BMP ligand expression could contribute to a high protein level of β-CATENIN in breast cancer cells. We demonstrated that downstream of BMP stimulation, SMAD4 is required to reduce β-CATENIN protein stability through the phosphorylation in MCF7 and T47D cells. Consequently, BMP stimulation reduces β-CATENIN levels and prevents its nuclear translocation and target gene expression in MCF7 cells. Conversely, BMP stimulation has no effect on β-CATENIN phosphorylation or stability in MDA-MB-231 and MDA-MB-468 cells. Likewise, SMAD4 modulation does not alter the response of those cells, indicating that SMAD4 alone is insufficient for BMP-induced β-CATENIN phosphorylation. While our data suggest that considering BMP activity may serve as a prognostic marker for understanding β-CATENIN accumulation risk, further investigation is needed to elucidate the differential responsiveness of breast cancer cell lines.
Full article
(This article belongs to the Special Issue Alterations to Signalling Pathways in Cancer Cells 2024)
►▼
Show Figures
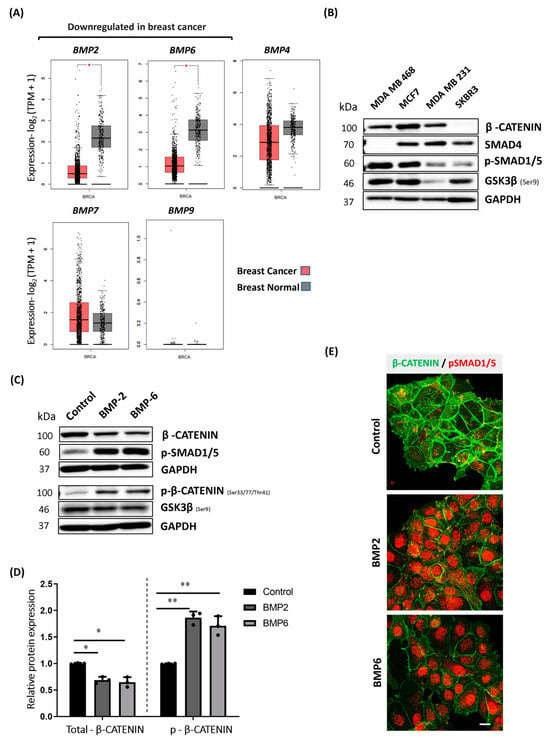
Figure 1
Open AccessArticle
Exogenous Melatonin Enhances Dihydrochalcone Accumulation in Lithocarpus litseifolius Leaves via Regulating Hormonal Crosstalk and Transcriptional Profiling
by
Wenlong Zhang, Yuqi Sun, Hongfeng Wang, Mingfeng Xu, Chunmei He, Congcong Wang, Yongli Yu, Zongshen Zhang and Lingye Su
Int. J. Mol. Sci. 2024, 25(9), 4592; https://doi.org/10.3390/ijms25094592 (registering DOI) - 23 Apr 2024
Abstract
Dihydrochalcones (DHCs) constitute a specific class of flavonoids widely known for their various health-related advantages. Melatonin (MLT) has received attention worldwide as a master regulator in plants, but its roles in DHC accumulation remain unclear. Herein, the elicitation impacts of MLT on DHC
[...] Read more.
Dihydrochalcones (DHCs) constitute a specific class of flavonoids widely known for their various health-related advantages. Melatonin (MLT) has received attention worldwide as a master regulator in plants, but its roles in DHC accumulation remain unclear. Herein, the elicitation impacts of MLT on DHC biosynthesis were examined in Lithocarpus litseifolius, a valuable medicinal plant famous for its sweet flavor and anti-diabetes effect. Compared to the control, the foliar application of MLT significantly increased total flavonoid and DHC (phlorizin, trilobatin, and phloretin) levels in L. litseifolius leaves, especially when 100 μM MLT was utilized for 14 days. Moreover, antioxidant enzyme activities were boosted after MLT treatments, resulting in a decrease in the levels of intracellular reactive oxygen species. Remarkably, MLT triggered the biosynthesis of numerous phytohormones linked to secondary metabolism (salicylic acid, methyl jasmonic acid (MeJA), and ethylene), while reducing free JA contents in L. litseifolius. Additionally, the flavonoid biosynthetic enzyme activities were enhanced by the MLT in leaves. Multiple differentially expressed genes (DEGs) in RNA-seq might play a crucial role in MLT-elicited pathways, particularly those associated with the antioxidant system (SOD, CAT, and POD), transcription factor regulation (MYBs and bHLHs), and DHC metabolism (4CL, C4H, UGT71K1, and UGT88A1). As a result, MLT enhanced DHC accumulation in L. litseifolius leaves, primarily by modulating the antioxidant activity and co-regulating the physiological, hormonal, and transcriptional pathways of DHC metabolism.
Full article
(This article belongs to the Special Issue Boosting the Production of Bioactive Compounds: Biotechnology and Metabolic Engineering of Medicinal Plants)
►▼
Show Figures
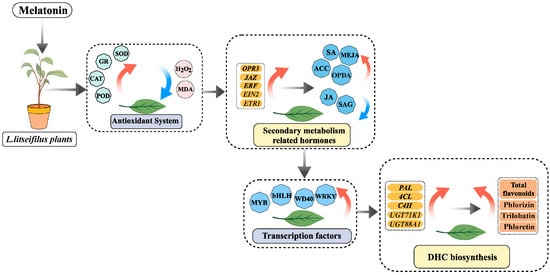
Figure 1
Open AccessArticle
Synthesis, In Silico and Kinetics Evaluation of N-(β-d-glucopyranosyl)-2-arylimidazole-4(5)-carboxamides and N-(β-d-glucopyranosyl)-4(5)-arylimidazole-2-carboxamides as Glycogen Phosphorylase Inhibitors
by
Levente Homolya, Rachel T. Mathomes, Luca Varga, Tibor Docsa, László Juhász, Joseph M. Hayes and László Somsák
Int. J. Mol. Sci. 2024, 25(9), 4591; https://doi.org/10.3390/ijms25094591 (registering DOI) - 23 Apr 2024
Abstract
Recently studied N-(β-d-glucopyranosyl)-3-aryl-1,2,4-triazole-5-carboxamides have proven to be low micromolar inhibitors of glycogen phosphorylase (GP), a validated target for the treatment of type 2 diabetes mellitus. Since in other settings, the bioisosteric replacement of the 1,2,4-triazole moiety with imidazole resulted
[...] Read more.
Recently studied N-(β-d-glucopyranosyl)-3-aryl-1,2,4-triazole-5-carboxamides have proven to be low micromolar inhibitors of glycogen phosphorylase (GP), a validated target for the treatment of type 2 diabetes mellitus. Since in other settings, the bioisosteric replacement of the 1,2,4-triazole moiety with imidazole resulted in significantly more efficient GP inhibitors, in silico calculations using Glide molecular docking along with unbound state DFT calculations were performed on N-(β-d-glucopyranosyl)-arylimidazole-carboxamides, revealing their potential for strong GP inhibition. The syntheses of the target compounds involved the formation of an amide bond between per-O-acetylated β-d-glucopyranosylamine and the corresponding arylimidazole-carboxylic acids. Kinetics experiments on rabbit muscle GPb revealed low micromolar inhibitors, with the best inhibition constants (Kis) of ~3–4 µM obtained for 1- and 2-naphthyl-substituted N-(β-d-glucopyranosyl)-imidazolecarboxamides, 2b–c. The predicted protein–ligand interactions responsible for the observed potencies are discussed and will facilitate the structure-based design of other inhibitors targeting this important therapeutic target. Meanwhile, the importance of the careful consideration of ligand tautomeric states in binding calculations is highlighted, with the usefulness of DFT calculations in this regard proposed.
Full article
(This article belongs to the Special Issue 21st Century Enzymology: Further Advances in Identification and Characterization of Enzymes)
►▼
Show Figures
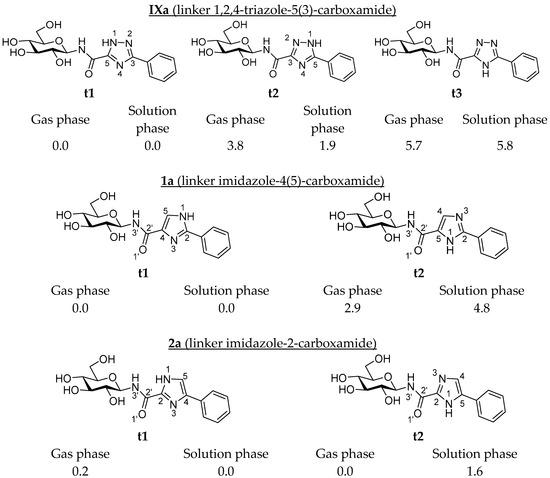
Figure 1
Open AccessArticle
Starch-Based Polysaccharide Systems with Bioactive Substances: Physicochemical and Wettability Characteristics
by
Agnieszka Ewa Wiącek and Anna Furmaniuk
Int. J. Mol. Sci. 2024, 25(9), 4590; https://doi.org/10.3390/ijms25094590 (registering DOI) - 23 Apr 2024
Abstract
Polysaccharide-based systems have very good emulsifying and stabilizing properties, and starch plays a leading role. Their modifications should add new quality features to the product to such an extent that preserves the structure-forming properties of native starch. The aim of this manuscript was
[...] Read more.
Polysaccharide-based systems have very good emulsifying and stabilizing properties, and starch plays a leading role. Their modifications should add new quality features to the product to such an extent that preserves the structure-forming properties of native starch. The aim of this manuscript was to examine the physicochemical characteristics of the combinations of starch with phospholipids or lysozymes and determine the effect of starch modification (surface hydrophobization or biological additives) and preparation temperature (before and after gelatinization). Changes in electrokinetic potential (zeta), effective diameter, and size distribution as a function of time were analyzed using the dynamic light scattering and microelectrophoresis techniques. The wettability of starch-coated glass plates before and after modification was checked by the advancing and receding contact angle measurements, as well as the angle hysteresis, using the settle drop method as a complement to profilometry and FTIR. It can be generalized that starch dispersions are more stable than analogous n-alkane/starch emulsions at room and physiological temperatures. On the other hand, the contact angle hysteresis values usually decrease with temperature increase, pointing to a more homogeneous surface, and the hydrophobization effect decreases vs. the thickness of the substrate. Surface hydrophobization of starch carried out using an n-alkane film does not change its bulk properties and leads to improvement of its mechanical and functional properties. The obtained specific starch-based hybrid systems, characterized in detail by switchable wettability, give the possibility to determine the energetic state of the starch surface and understand the strength and specificity of interactions with substances of different polarities in biological processes and their applicability for multidirectional use.
Full article
(This article belongs to the Special Issue New Perspectives of Colloids for Biological Applications)
►▼
Show Figures
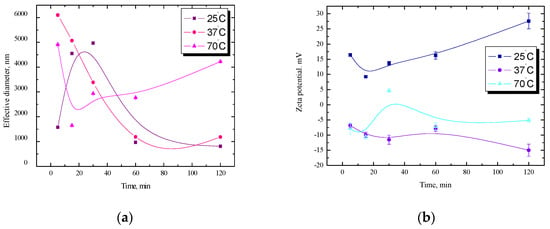
Figure 1
Open AccessArticle
Modification of Seurat v4 for the Development of a Phase Assignment Tool Able to Distinguish between G2 and Mitotic Cells
by
Steven Watson, Harry Porter, Ian Sudbery and Ruth Thompson
Int. J. Mol. Sci. 2024, 25(9), 4589; https://doi.org/10.3390/ijms25094589 (registering DOI) - 23 Apr 2024
Abstract
Single-cell RNA sequencing (scRNAseq) is a rapidly advancing field enabling the characterisation of heterogeneous gene expression profiles within a population. The cell cycle phase is a major contributor to gene expression variance between cells and computational analysis tools have been developed to assign
[...] Read more.
Single-cell RNA sequencing (scRNAseq) is a rapidly advancing field enabling the characterisation of heterogeneous gene expression profiles within a population. The cell cycle phase is a major contributor to gene expression variance between cells and computational analysis tools have been developed to assign cell cycle phases to cells within scRNAseq datasets. Whilst these tools can be extremely useful, all have the drawback that they classify cells as only G1, S or G2/M. Existing discrete cell phase assignment tools are unable to differentiate between G2 and M and continuous-phase-assignment tools are unable to identify a region corresponding specifically to mitosis in a pseudo-timeline for continuous assignment along the cell cycle. In this study, bulk RNA sequencing was used to identify differentially expressed genes between mitotic and interphase cells isolated based on phospho-histone H3 expression using fluorescence-activated cell sorting. These gene lists were used to develop a methodology which can distinguish G2 and M phase cells in scRNAseq datasets. The phase assignment tools present in Seurat were modified to allow for cell cycle phase assignment of all stages of the cell cycle to identify a mitotic-specific cell population.
Full article
(This article belongs to the Special Issue Latest Progress in DNA Damage and DNA Repair)
►▼
Show Figures
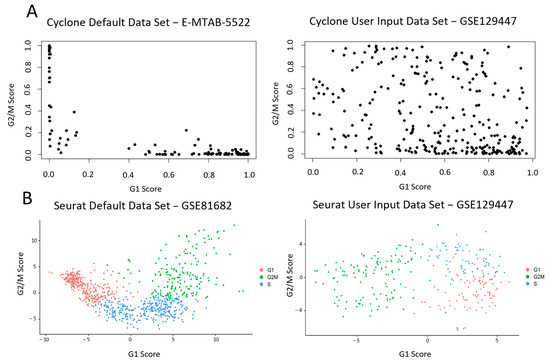
Figure 1
Open AccessArticle
Elucidating the Role of circTIAM1 in Guangling Large-Tailed Sheep Adipocyte Proliferation and Differentiation via the miR-485-3p/PLCB1 Pathway
by
Yu Liang, Bishi Zhao, Yan Shen, Miao Peng, Liying Qiao, Jianhua Liu, Yangyang Pan, Kaijie Yang and Wenzhong Liu
Int. J. Mol. Sci. 2024, 25(9), 4588; https://doi.org/10.3390/ijms25094588 (registering DOI) - 23 Apr 2024
Abstract
Fat tissue—a vital energy storage organ—is intricately regulated by various factors, including circular RNA, which plays a significant role in modulating fat development and lipid metabolism. Therefore, this study aims to clarify the regulatory mechanism of sheep adipocyte proliferation and differentiation by investigating
[...] Read more.
Fat tissue—a vital energy storage organ—is intricately regulated by various factors, including circular RNA, which plays a significant role in modulating fat development and lipid metabolism. Therefore, this study aims to clarify the regulatory mechanism of sheep adipocyte proliferation and differentiation by investigating the involvement of circTIAM1, miR-485-3p, and its target gene PLCB1. Through previous sequencing data, circTIAM1 was identified in sheep adipocytes, with its circularization mechanism elucidated, confirming its cytoplasmic localization. Experimental evidence from RNase R treatment and transcription inhibitors highlighted that circTIAM1 is more stable than linear RNA. Additionally, circTIAM1 promoted sheep adipocyte proliferation and differentiation. Furthermore, bioinformatic analysis demonstrated a robust interaction between miR-485-3p and circTIAM1. Further experiments revealed that miR-485-3p inhibits fat cell proliferation and differentiation by inhibiting PLCB1, with circTIAM1 alleviating the inhibitory effect via competitive binding. In summary, our findings elucidate the mechanism through which circTIAM1 regulates Guangling Large-Tailed sheep adipocyte proliferation and differentiation via the miR-485-3p–PLCB1 pathway, offering a novel perspective for further exploring fat metabolism regulation.
Full article
(This article belongs to the Section Molecular Biology)
►▼
Show Figures

Figure 1
Open AccessArticle
Halogen Bond via an Electrophilic π-Hole on Halogen in Molecules: Does It Exist?
by
Pradeep R. Varadwaj
Int. J. Mol. Sci. 2024, 25(9), 4587; https://doi.org/10.3390/ijms25094587 (registering DOI) - 23 Apr 2024
Abstract
This study reveals a new non-covalent interaction called a π-hole halogen bond, which is directional and potentially non-linear compared to its sister analog (σ-hole halogen bond). A π-hole is shown here to be observed on the surface of halogen in halogenated molecules, which
[...] Read more.
This study reveals a new non-covalent interaction called a π-hole halogen bond, which is directional and potentially non-linear compared to its sister analog (σ-hole halogen bond). A π-hole is shown here to be observed on the surface of halogen in halogenated molecules, which can be tempered to display the aptness to form a π-hole halogen bond with a series of electron density-rich sites (Lewis bases) hosted individually by 32 other partner molecules. The [MP2/aug-cc-pVTZ] level characteristics of the π-hole halogen bonds in 33 binary complexes obtained from the charge density approaches (quantum theory of intramolecular atoms, molecular electrostatic surface potential, independent gradient model (IGM-δginter)), intermolecular geometries and energies, and second-order hyperconjugative charge transfer analyses are discussed, which are similar to other non-covalent interactions. That a π-hole can be observed on halogen in halogenated molecules is substantiated by experimentally reported crystals documented in the Cambridge Crystal Structure Database. The importance of the π-hole halogen bond in the design and growth of chemical systems in synthetic chemistry, crystallography, and crystal engineering is yet to be fully explicated.
Full article
(This article belongs to the Special Issue Noncovalent Interactions: New Developments in Experiment and Theory)
►▼
Show Figures
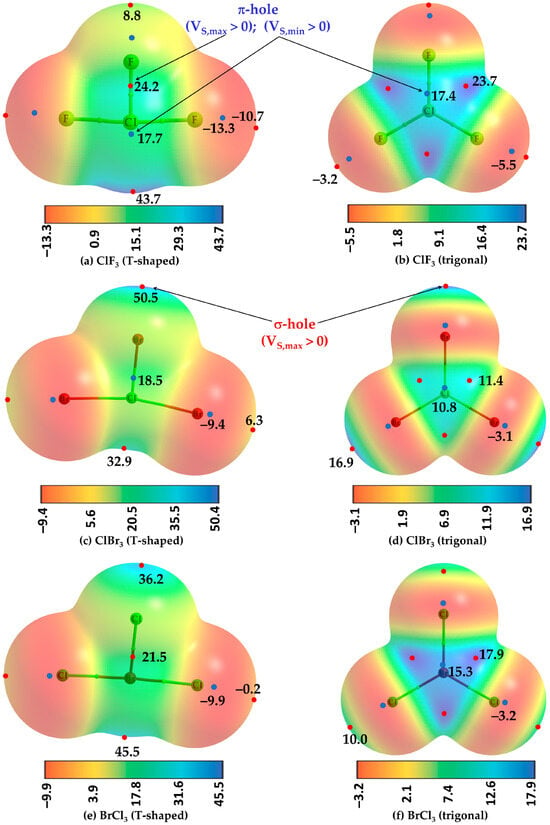
Figure 1
Open AccessEditorial
Special Issue “Physiology and Pathophysiology of Placenta 2.0”
by
Giovanni Tossetta
Int. J. Mol. Sci. 2024, 25(9), 4586; https://doi.org/10.3390/ijms25094586 (registering DOI) - 23 Apr 2024
Abstract
We are pleased to present this Special Issue of the International Journal of Molecular Sciences, entitled “Physiology and Pathophysiology of Placenta 2 [...]
Full article
(This article belongs to the Special Issue Physiology and Pathophysiology of Placenta 2.0)
Open AccessArticle
ZmNAC17 Regulates Mesocotyl Elongation by Mediating Auxin and ROS Biosynthetic Pathways in Maize
by
Ran Yang, Kangshi Li, Ming Wang, Meng Sun, Qiuhua Li, Liping Chen, Feng Xiao, Zhenlong Zhang, Haiyan Zhang, Fuchao Jiao and Jingtang Chen
Int. J. Mol. Sci. 2024, 25(9), 4585; https://doi.org/10.3390/ijms25094585 (registering DOI) - 23 Apr 2024
Abstract
The mesocotyl is of great significance in seedling emergence and in responding to biotic and abiotic stress in maize. The NAM, ATAF, and CUC2 (NAC) transcription factor family plays an important role in maize growth and development; however, its function in the elongation
[...] Read more.
The mesocotyl is of great significance in seedling emergence and in responding to biotic and abiotic stress in maize. The NAM, ATAF, and CUC2 (NAC) transcription factor family plays an important role in maize growth and development; however, its function in the elongation of the maize mesocotyl is still unclear. In this study, we found that the mesocotyl length in zmnac17 loss-of-function mutants was lower than that in the B73 wild type. By using transcriptomic sequencing technology, we identified 444 differentially expressed genes (DEGs) between zmnac17-1 and B73, which were mainly enriched in the “tryptophan metabolism” and “antioxidant activity” pathways. Compared with the control, the zmnac17-1 mutants exhibited a decrease in the content of indole acetic acid (IAA) and an increase in the content of reactive oxygen species (ROS). Our results provide preliminary evidence that ZmNAC17 regulates the elongation of the maize mesocotyl.
Full article
(This article belongs to the Special Issue Plant Hormone Signaling)
►▼
Show Figures

Figure 1
Open AccessArticle
The Impact of Normobaric Hypoxia and Intermittent Hypoxic Training on Cardiac Biomarkers in Endurance Athletes: A Pilot Study
by
Jakub Goliniewski, Miłosz Czuba, Kamila Płoszczyca, Małgorzata Chalimoniuk, Robert Gajda, Adam Niemaszyk, Katarzyna Kaczmarczyk and Józef Langfort
Int. J. Mol. Sci. 2024, 25(9), 4584; https://doi.org/10.3390/ijms25094584 (registering DOI) - 23 Apr 2024
Abstract
This study explores the effects of normobaric hypoxia and intermittent hypoxic training (IHT) on the physiological condition of the cardiac muscle in swimmers. Hypoxia has been reported to elicit both beneficial and adverse changes in the cardiovascular system, but its impact on the
[...] Read more.
This study explores the effects of normobaric hypoxia and intermittent hypoxic training (IHT) on the physiological condition of the cardiac muscle in swimmers. Hypoxia has been reported to elicit both beneficial and adverse changes in the cardiovascular system, but its impact on the myocardium during acute exercise and altitude/hypoxic training remains less understood. We aimed to determine how a single bout of intense interval exercise and a four-week period of high-intensity endurance training under normobaric hypoxia affect cardiac marker activity in swimmers. Sixteen young male swimmers were divided into two groups: one undergoing training in hypoxia and the other in normoxia. Cardiac markers, including troponin I and T (cTnI and cTnT), heart-type fatty acid-binding protein (H-FABP), creatine kinase-MB isoenzyme (CK-MB), and myoglobin (Mb), were analyzed to assess the myocardium’s response. We found no significant differences in the physiological response of the cardiac muscle to intense physical exertion between hypoxia and normoxia. Four weeks of IHT did not alter the resting levels of cTnT, cTnI, and H-FABP, but it resulted in a noteworthy decrease in the resting concentration of CK-MB, suggesting enhanced cardiac muscle adaptation to exercise. In contrast, a reduction in resting Mb levels was observed in the control group training in normoxia. These findings suggest that IHT at moderate altitudes does not adversely affect cardiac muscle condition and may support cardiac muscle adaptation, affirming the safety and efficacy of IHT as a training method for athletes.
Full article
(This article belongs to the Section Molecular Pathology, Diagnostics, and Therapeutics)
►▼
Show Figures
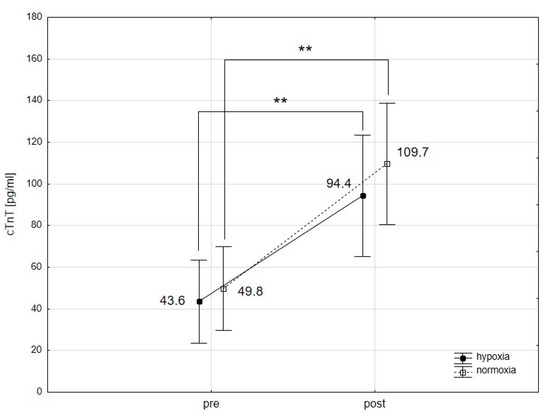
Figure 1
Open AccessArticle
Enhanced ALOX12 Gene Expression Predicts Therapeutic Susceptibility to 5-Azacytidine in Patients with Myelodysplastic Syndromes
by
Taichi Matsumoto, Yuichi Murakami, Nao Yoshida-Sakai, Daisuke Katsuchi, Kuon Kanazawa, Takashi Okamura, Yutaka Imamura, Mayumi Ono and Michihiko Kuwano
Int. J. Mol. Sci. 2024, 25(9), 4583; https://doi.org/10.3390/ijms25094583 (registering DOI) - 23 Apr 2024
Abstract
5-azacytidine (AZA), a representative DNA-demethylating drug, has been widely used to treat myelodysplastic syndromes (MDS). However, it remains unclear whether AZA’s DNA demethylation of any specific gene is correlated with clinical responses to AZA. In this study, we investigated genes that could contribute
[...] Read more.
5-azacytidine (AZA), a representative DNA-demethylating drug, has been widely used to treat myelodysplastic syndromes (MDS). However, it remains unclear whether AZA’s DNA demethylation of any specific gene is correlated with clinical responses to AZA. In this study, we investigated genes that could contribute to the development of evidence-based epigenetic therapeutics with AZA. A DNA microarray identified that AZA specifically upregulated the expression of 438 genes in AZA-sensitive MDS-L cells but not in AZA-resistant counterpart MDS-L/CDA cells. Of these 438 genes, the ALOX12 gene was hypermethylated in MDS-L cells but not in MDS-L/CDA cells. In addition, we further found that (1) the ALOX12 gene was hypermethylated in patients with MDS compared to healthy controls; (2) MDS classes with excess blasts showed a relatively lower expression of ALOX12 than other classes; (3) a lower expression of ALOX12 correlated with higher bone marrow blasts and a shorter survival in patients with MDS; and (4) an increased ALOX12 expression after AZA treatment was associated with a favorable response to AZA treatment. Taking these factors together, an enhanced expression of the ALOX12 gene may predict favorable therapeutic responses to AZA therapy in MDS.
Full article
(This article belongs to the Special Issue Leukemia: Present and Future)
►▼
Show Figures
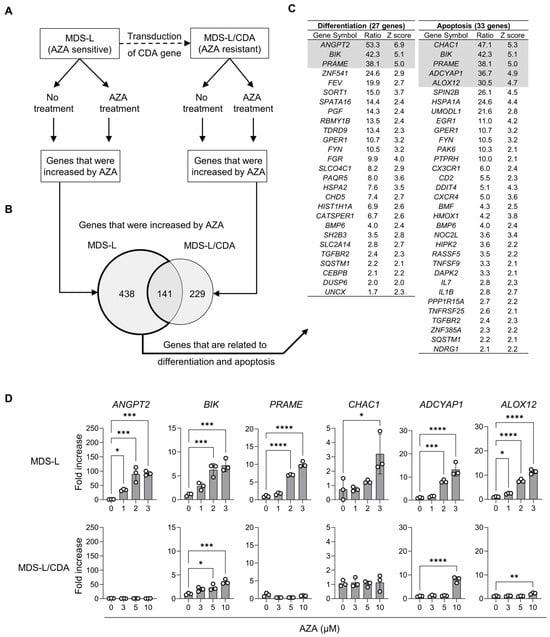
Figure 1

Journal Menu
► ▼ Journal Menu-
- IJMS Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal Browser-
arrow_forward_ios
Forthcoming issue
arrow_forward_ios Current issue - Vol. 25 (2024)
- Vol. 24 (2023)
- Vol. 23 (2022)
- Vol. 22 (2021)
- Vol. 21 (2020)
- Vol. 20 (2019)
- Vol. 19 (2018)
- Vol. 18 (2017)
- Vol. 17 (2016)
- Vol. 16 (2015)
- Vol. 15 (2014)
- Vol. 14 (2013)
- Vol. 13 (2012)
- Vol. 12 (2011)
- Vol. 11 (2010)
- Vol. 10 (2009)
- Vol. 9 (2008)
- Vol. 8 (2007)
- Vol. 7 (2006)
- Vol. 6 (2005)
- Vol. 5 (2004)
- Vol. 4 (2003)
- Vol. 3 (2002)
- Vol. 2 (2001)
- Vol. 1 (2000)
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomedicines, Cells, IJMS, Life, Oxygen
Oxidative Stress and Inflammation, 2nd Volume
Topic Editors: Mohamad Allaw, Ines Castangia, Maria Letizia Manca, Matteo Perra, Amparo NacherDeadline: 31 May 2024
Topic in
BioChem, Biomedicines, Biomolecules, IJMS, Metabolites, Molecules
Natural Products in Prevention and Therapy of Metabolic Syndrome
Topic Editors: Jianbo Wan, Ligen LinDeadline: 30 June 2024
Topic in
Cells, Diseases, Healthcare, IJMS, Vaccines
Inflammation: The Cause of all Diseases 2.0
Topic Editors: Vasso Apostolopoulos, Jack Feehan, Vivek P. ChavdaDeadline: 31 July 2024
Topic in
Biomedicines, CIMB, Endocrines, IJMS, JMP, Life, Reprod. Med.
Pathogenesis of Pregnancy-Related Complications 2.0
Topic Editors: Ilona Hromadnikova, Katerina KotlabovaDeadline: 31 August 2024

Conferences
Special Issues
Special Issue in
IJMS
Structural Dynamics of Macromolecules
Guest Editor: Ki Hyun NamDeadline: 25 April 2024
Special Issue in
IJMS
Monoclonal Antibodies and Their Functional Fragments in Research, Diagnosis and Therapy 3.0
Guest Editors: Menotti Ruvo, Annamaria SandomenicoDeadline: 30 April 2024
Special Issue in
IJMS
Pharmacogenetics and Personalized Medicine 3.0
Guest Editors: José A. Riancho, Gloria RavegniniDeadline: 20 May 2024
Special Issue in
IJMS
Molecular Mechanisms of Angiogenesis and Cancer
Guest Editor: Vijay AvinDeadline: 30 May 2024
Topical Collections
Topical Collection in
IJMS
Feature Papers in Bioactives and Nutraceuticals
Collection Editor: Maurizio Battino
Topical Collection in
IJMS
State-of-the-Art Molecular Microbiology in Poland
Collection Editors: Alicja Wegrzyn, Satish Raina
Topical Collection in
IJMS
Computational, Structural and Spectroscopic Studies of Enzyme Mechanisms, Inhibition and Dynamics
Collection Editor: Christo Christov