Journal Description
Viruses
Viruses
is a peer-reviewed, open access journal of virology, published monthly online by MDPI. The American Society for Virology (ASV), Spanish Society for Virology (SEV), Canadian Society for Virology (CSV), Italian Society for Virology (SIV-ISV), Australasian Virology Society (AVS) and others are affiliated with Viruses and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Virology) / CiteScore - Q1 (Infectious Diseases)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 13.8 days after submission; acceptance to publication is undertaken in 2.5 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Companion journals for Viruses include: COVID and Zoonotic Diseases.
Impact Factor:
4.7 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
Expanding the Scope of Adenoviral Vectors by Utilizing Novel Tools for Recombination and Vector Rescue
Viruses 2024, 16(5), 658; https://doi.org/10.3390/v16050658 - 23 Apr 2024
Abstract
Recombinant adenoviruses are widely used in clinical and laboratory applications. Despite the wide variety of available sero- and genotypes, only a fraction is utilized in vivo. As adenoviruses are a large group of viruses, displaying many different tropisms, immune epitopes, and replication characteristics,
[...] Read more.
Recombinant adenoviruses are widely used in clinical and laboratory applications. Despite the wide variety of available sero- and genotypes, only a fraction is utilized in vivo. As adenoviruses are a large group of viruses, displaying many different tropisms, immune epitopes, and replication characteristics, the merits of translating these natural benefits into vector applications are apparent. This translation, however, proves difficult, since while research has investigated the application of these viruses, there are no universally applicable rules in vector design for non-classical adenovirus types. In this paper, we describe a generalized workflow that allows vectorization, rescue, and cloning of all adenoviral species to enable the rapid development of new vector variants. We show this using human and simian adenoviruses, further modifying a selection of them to investigate their gene transfer potential and build potential vector candidates for future applications.
Full article
(This article belongs to the Special Issue 15th International Adenovirus Meeting)
Open AccessArticle
Wide Real-Life Data Support Reduced Sensitivity of Antigen Tests for Omicron SARS-CoV-2 Infections
by
Chiara Piubelli, Davide Treggiari, Denise Lavezzari, Michela Deiana, Klevia Dishnica, Emma Maria Sole Tosato, Cristina Mazzi, Paolo Cattaneo, Antonio Mori, Elena Pomari, Lavinia Nicolini, Martina Leonardi, Francesca Perandin, Fabio Formenti, Alejandro Giorgetti, Antonio Conti, Maria Rosaria Capobianchi, Federico Giovanni Gobbi and Concetta Castilletti
Viruses 2024, 16(5), 657; https://doi.org/10.3390/v16050657 - 23 Apr 2024
Abstract
With the continuous spread of new SARS-CoV-2 variants of concern (VOCs), the monitoring of diagnostic test performances is mandatory. We evaluated the changes in antigen diagnostic tests’ (ADTs) accuracy along the Delta to Omicron VOCs transition, exploring the N protein mutations possibly affecting
[...] Read more.
With the continuous spread of new SARS-CoV-2 variants of concern (VOCs), the monitoring of diagnostic test performances is mandatory. We evaluated the changes in antigen diagnostic tests’ (ADTs) accuracy along the Delta to Omicron VOCs transition, exploring the N protein mutations possibly affecting ADT sensitivity and assessing the best sampling site for the diagnosis of Omicron infections. In total, 5175 subjects were enrolled from 1 October 2021 to 15 July 2022. The inclusion criteria were SARS-CoV-2 ADT combined with a same-day RT-PCR swab test. For the sampling site analysis, 61 patients were prospectively recruited during the Omicron period for nasal and oral swab analyses by RT-PCR. Next-Generation Sequencing data were obtained to evaluate the different sublineages. Using RT-PCR as a reference, 387 subjects resulted in becoming infected and the overall sensitivity of the ADT decreased from 63% in the Delta period to 33% in the Omicron period. This decrease was highly statistically significant (p < 0.001), and no decrease in viral load was detected at the RNA level. The nasal site presented a significantly higher viral load than the oral site during the Omicron wave. The reduced detection rate of Omicron infections by ADT should be considered in the global testing strategy to preserve accurate diagnoses across the changing SARS-CoV-2 variants.
Full article
(This article belongs to the Section Coronaviruses)
►▼
Show Figures
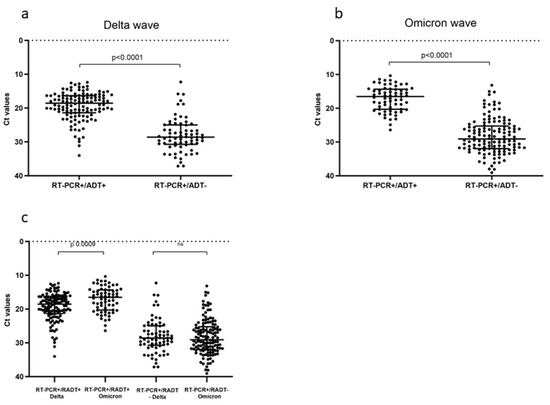
Figure 1
Open AccessArticle
The Changing Detection Rate of Respiratory Syncytial Virus in Adults in Western Australia between 2017 and 2023
by
David A. Foley, Cara A. Minney-Smith, Andrew Tjea, Mark P. Nicol, Avram Levy, Hannah C. Moore and Christopher C. Blyth
Viruses 2024, 16(5), 656; https://doi.org/10.3390/v16050656 - 23 Apr 2024
Abstract
The incidence of respiratory syncytial virus (RSV) in adults is inadequately defined and the impact of SARS-CoV-2-related non-pharmaceutical interventions (NPIs) is underexplored. Using laboratory data, we described the detection rate of RSV in adults ≥16 years in Western Australia (WA) between 2017 and
[...] Read more.
The incidence of respiratory syncytial virus (RSV) in adults is inadequately defined and the impact of SARS-CoV-2-related non-pharmaceutical interventions (NPIs) is underexplored. Using laboratory data, we described the detection rate of RSV in adults ≥16 years in Western Australia (WA) between 2017 and 2023. With the exception of 2020, RSV detections rose annually between 2017 and 2023, reaching 50.7 per 100,000 in 2023 (95% confidence interval [CI], 47.9–53.8). RSV testing expanded considerably across the study period, with the testing in 2023 more than five times the 2017 total. The detection rate was highest in adults ≥60 years between 2017 and 2019, particularly those ≥75 years. Following 2020, the detections in all age groups increased, with the highest detection rate in 2023 in those ≥75-years (199.5 per 100,000; 95% CI, 180.5–220). NPIs significantly impacted RSV seasonality; the preceding winter pattern was disrupted, resulting in an absent 2020 winter season and two major summer seasons in 2020/21 and 2021/22. The RSV season began to realign in 2022, reverting to a winter seasonal pattern in 2023 and the largest season in the study period. Ongoing surveillance will be required to understand the stability of these increases and to delineate the impact of new immunisation strategies.
Full article
(This article belongs to the Special Issue Impact of Pandemic Measures on the Epidemiology and Seasonality of Other Viruses)
►▼
Show Figures
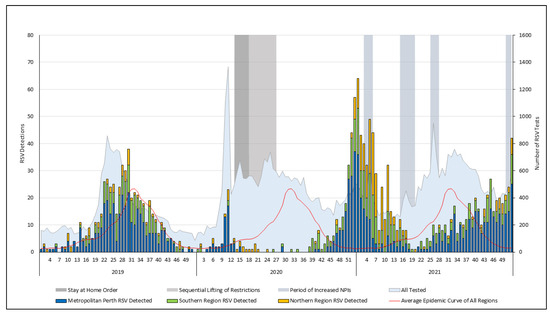
Figure 1
Open AccessArticle
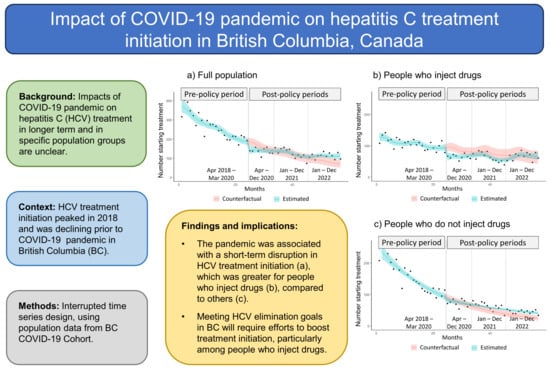
Impact of the COVID-19 Pandemic on Hepatitis C Treatment Initiation in British Columbia, Canada: An Interrupted Time Series Study
by
Richard L. Morrow, Mawuena Binka, Julia Li, Mike Irvine, Sofia R. Bartlett, Stanley Wong, Dahn Jeong, Jean Damascene Makuza, Jason Wong, Amanda Yu, Mel Krajden and Naveed Zafar Janjua
Viruses 2024, 16(5), 655; https://doi.org/10.3390/v16050655 - 23 Apr 2024
Abstract
We investigated the impacts of the COVID-19 pandemic on hepatitis C (HCV) treatment initiation, including by birth cohort and injection drug use status, in British Columbia (BC), Canada. Using population data from the BC COVID-19 Cohort, we conducted interrupted time series analyses, estimating
[...] Read more.
We investigated the impacts of the COVID-19 pandemic on hepatitis C (HCV) treatment initiation, including by birth cohort and injection drug use status, in British Columbia (BC), Canada. Using population data from the BC COVID-19 Cohort, we conducted interrupted time series analyses, estimating changes in HCV treatment initiation following the introduction of pandemic-related policies in March 2020. The study included a pre-policy period (April 2018 to March 2020) and three follow-up periods (April to December 2020, January to December 2021, and January to December 2022). The level of HCV treatment initiation decreased by 26% in April 2020 (rate ratio 0.74, 95% confidence interval [CI] 0.60 to 0.91). Overall, no statistically significant difference in HCV treatment initiation occurred over the 2020 and 2021 post-policy periods, and an increase of 34.4% (95% CI 0.6 to 75.8) occurred in 2022 (equating to 321 additional people initiating treatment), relative to expectation. Decreases in HCV treatment initiation occurred in 2020 for people born between 1965 and 1974 (25.5%) and people who inject drugs (24.5%), relative to expectation. In summary, the pandemic was associated with short-term disruptions in HCV treatment initiation in BC, which were greater for people born 1965 to 1974 and people who inject drugs.
Full article
(This article belongs to the Special Issue Cascade of Care for HIV and Hepatitis)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Epidemiological Features of Human Norovirus Genotypes before and after COVID-19 Countermeasures in Osaka, Japan
by
Tatsuya Shirai, Juthamas Phadungsombat, Yumi Ushikai, Kunihito Yoshikaie, Tatsuo Shioda and Naomi Sakon
Viruses 2024, 16(4), 654; https://doi.org/10.3390/v16040654 - 22 Apr 2024
Abstract
We investigated the molecular epidemiology of human norovirus (HuNoV) in all age groups using samples from April 2019 to March 2023, before and after the COVID-19 countermeasures were implemented. GII.2[P16] and GII.4[P31], the prevalent strains in Japan before COVID-19 countermeasures, remained prevalent during
[...] Read more.
We investigated the molecular epidemiology of human norovirus (HuNoV) in all age groups using samples from April 2019 to March 2023, before and after the COVID-19 countermeasures were implemented. GII.2[P16] and GII.4[P31], the prevalent strains in Japan before COVID-19 countermeasures, remained prevalent during the COVID-19 pandemic, except from April to November 2020; in 2021, the prevalence of GII.2[P16] increased among children. Furthermore, there was an increase in the prevalence of GII.4[P16] after December 2022. Phylogenetic analysis of GII.P31 RdRp showed that some strains detected in 2022 belonged to a different cluster of other strains obtained during the present study period, suggesting that HuNoV strains will evolve differently even if they have the same type of RdRp. An analysis of the amino acid sequence of VP1 showed that some antigenic sites of GII.4[P16] were different from those of GII.4[P31]. The present study showed high infectivity of HuNoV despite the COVID-19 countermeasures and revealed changes in the prevalent genotypes and mutations of each genotype. In the future, we will investigate whether GII.4[P16] becomes more prevalent, providing new insights by comparing the new data with those analyzed in the present study.
Full article
(This article belongs to the Special Issue Viral Genetic Variation)
►▼
Show Figures

Figure 1
Open AccessArticle
Delving into the Aftermath of a Disease-Associated Near-Extinction Event: A Five-Year Study of a Serpentovirus (Nidovirus) in a Critically Endangered Turtle Population
by
Kate Parrish, Peter Kirkland, Paul Horwood, Bruce Chessman, Shane Ruming, Gerry McGilvray, Karrie Rose, Jane Hall and Lee Skerratt
Viruses 2024, 16(4), 653; https://doi.org/10.3390/v16040653 - 22 Apr 2024
Abstract
Bellinger River virus (BRV) is a serpentovirus (nidovirus) that was likely responsible for the catastrophic mortality of the Australian freshwater turtle Myuchelys georgesi in February 2015. From November 2015 to November 2020, swabs were collected from turtles during repeated river surveys to estimate
[...] Read more.
Bellinger River virus (BRV) is a serpentovirus (nidovirus) that was likely responsible for the catastrophic mortality of the Australian freshwater turtle Myuchelys georgesi in February 2015. From November 2015 to November 2020, swabs were collected from turtles during repeated river surveys to estimate the prevalence of BRV RNA, identify risk factors associated with BRV infection, and refine sample collection. BRV RNA prevalence at first capture was significantly higher in M. georgesi (10.8%) than in a coexisting turtle, Emydura macquarii (1.0%). For M. georgesi, various risk factors were identified depending on the analysis method, but a positive BRV result was consistently associated with a larger body size. All turtles were asymptomatic when sampled and conjunctival swabs were inferred to be optimal for ongoing monitoring. Although the absence of disease and recent BRV detections suggests a reduced ongoing threat, the potential for the virus to persist in an endemic focus or resurge in cyclical epidemics cannot be excluded. Therefore, BRV is an ongoing potential threat to the conservation of M. georgesi, and strict adherence to biosecurity principles is essential to minimise the risk of reintroduction or spread of BRV or other pathogens.
Full article
(This article belongs to the Special Issue Identifying and Characterizing Viral Infections in Reptiles, Amphibians, Fish and Other Aquatic Species)
►▼
Show Figures
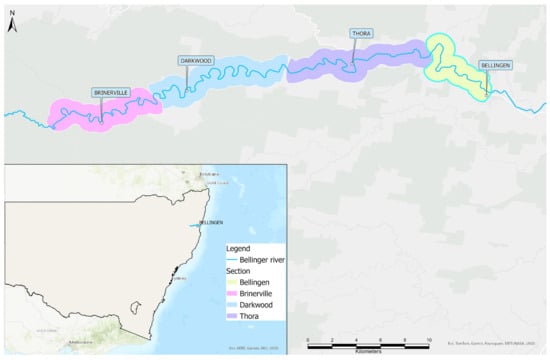
Figure 1
Open AccessArticle
Heterologous Exchanges of Glycoprotein and Non-Virion Protein in Novirhabdoviruses: Assessment of Virulence in Yellow Perch (Perca flavescens) and Rainbow Trout (Oncorhynchus mykiss)
by
Vikram N. Vakharia, Arun Ammayappan, Shamila Yusuff, Tarin M. Tesfaye and Gael Kurath
Viruses 2024, 16(4), 652; https://doi.org/10.3390/v16040652 - 22 Apr 2024
Abstract
Infectious hematopoietic necrosis virus (IHNV) and viral hemorrhagic septicemia virus (VHSV) are rhabdoviruses in two different species belonging to the Novirhabdovirus genus. IHNV has a narrow host range restricted to trout and salmon species, and viruses in the M genogroup of IHNV have
[...] Read more.
Infectious hematopoietic necrosis virus (IHNV) and viral hemorrhagic septicemia virus (VHSV) are rhabdoviruses in two different species belonging to the Novirhabdovirus genus. IHNV has a narrow host range restricted to trout and salmon species, and viruses in the M genogroup of IHNV have high virulence in rainbow trout (Oncorhynchus mykiss). In contrast, the VHSV genotype IVb that invaded the Great Lakes in the United States has a broad host range, with high virulence in yellow perch (Perca flavescens), but not in rainbow trout. By using reverse-genetic systems of IHNV-M and VHSV-IVb strains, we generated six IHNV:VHSV chimeric viruses in which the glycoprotein (G), non-virion-protein (NV), or both G and NV genes of IHNV-M were replaced with the analogous genes from VHSV-IVb, and vice versa. These chimeric viruses were used to challenge groups of rainbow trout and yellow perch. The parental recombinants rIHNV-M and rVHSV-IVb were highly virulent in rainbow trout and yellow perch, respectively. Parental rIHNV-M was avirulent in yellow perch, and chimeric rIHNV carrying G, NV, or G and NV genes from VHSV-IVb remained low in virulence in yellow perch. Similarly, the parental rVHSV-IVb exhibited low virulence in rainbow trout, and chimeric rVHSV with substituted G, NV, or G and NV genes from IHNV-M remained avirulent in rainbow trout. Thus, the G and NV genes of either virus were not sufficient to confer high host-specific virulence when exchanged into a heterologous species genome. Some exchanges of G and/or NV genes caused a loss of host-specific virulence, providing insights into possible roles in viral virulence or fitness, and interactions between viral proteins.
Full article
(This article belongs to the Special Issue The World of Rhabdoviruses)
►▼
Show Figures
Figure 1
Open AccessArticle
Why Certain Repurposed Drugs Are Unlikely to Be Effective Antivirals to Treat SARS-CoV-2 Infections
by
Selwyn J. Hurwitz, Ramyani De, Julia C. LeCher, Jessica A. Downs-Bowen, Shu Ling Goh, Keivan Zandi, Tamara McBrayer, Franck Amblard, Dharmeshkumar Patel, James J. Kohler, Manoj Bhasin, Brian S. Dobosh, Vikas Sukhatme, Rabindra M. Tirouvanziam and Raymond F. Schinazi
Viruses 2024, 16(4), 651; https://doi.org/10.3390/v16040651 - 22 Apr 2024
Abstract
Most repurposed drugs have proved ineffective for treating COVID-19. We evaluated median effective and toxic concentrations (EC50, CC50) of 49 drugs, mostly from previous clinical trials, in Vero cells. Ratios of reported unbound peak plasma concentrations, (Cmax)/EC
[...] Read more.
Most repurposed drugs have proved ineffective for treating COVID-19. We evaluated median effective and toxic concentrations (EC50, CC50) of 49 drugs, mostly from previous clinical trials, in Vero cells. Ratios of reported unbound peak plasma concentrations, (Cmax)/EC50, were used to predict the potential in vivo efficacy. The 20 drugs with the highest ratios were retested in human Calu-3 and Caco-2 cells, and their CC50 was determined in an expanded panel of cell lines. Many of the 20 drugs with the highest ratios were inactive in human Calu-3 and Caco-2 cells. Antivirals effective in controlled clinical trials had unbound Cmax/EC50 ≥ 6.8 in Calu-3 or Caco-2 cells. EC50 of nucleoside analogs were cell dependent. This approach and earlier availability of more relevant cultures could have reduced the number of unwarranted clinical trials.
Full article
(This article belongs to the Special Issue Advances in Antiviral Agents against SARS-CoV-2 and Its Variants 2nd Edition)
►▼
Show Figures

Graphical abstract
Open AccessCommunication
Association of the IFNG +874T/A Polymorphism with Symptomatic COVID-19 Susceptibility
by
Kevin Matheus Lima de Sarges, Flávia Póvoa da Costa, Erika Ferreira dos Santos, Marcos Henrique Damasceno Cantanhede, Rosilene da Silva, Adriana de Oliveira Lameira Veríssimo, Maria de Nazaré do Socorro de Almeida Viana, Fabíola Brasil Barbosa Rodrigues, Mauro de Meira Leite, Maria Karoliny da Silva Torres, Christiane Bentes da Silva, Mioni Thieli Figueiredo Magalhães de Brito, Andréa Luciana Soares da Silva, Daniele Freitas Henriques, Izaura Maria Vieira Cayres Vallinoto, Giselle Maria Rachid Viana, Maria Alice Freitas Queiroz, Antonio Carlos Rosário Vallinoto and Eduardo José Melo dos Santos
Viruses 2024, 16(4), 650; https://doi.org/10.3390/v16040650 - 22 Apr 2024
Abstract
Tumor necrosis factor (TNF) and interferon-gamma (IFNγ) are important inflammatory mediators in the development of cytokine storm syndrome (CSS). Single nucleotide polymorphisms (SNPs) regulate the expression of these cytokines, making host genetics a key factor in the prognosis of COVID-19. In this study,
[...] Read more.
Tumor necrosis factor (TNF) and interferon-gamma (IFNγ) are important inflammatory mediators in the development of cytokine storm syndrome (CSS). Single nucleotide polymorphisms (SNPs) regulate the expression of these cytokines, making host genetics a key factor in the prognosis of COVID-19. In this study, we investigated the associations of the TNF -308G/A and IFNG +874T/A polymorphisms with COVID-19. We analyzed the frequencies of the two polymorphisms in the control groups (CG: TNF -308G/A, n = 497; IFNG +874T/A, n = 397), a group of patients with COVID-19 (CoV, n = 222) and among the subgroups of patients with nonsevere (n = 150) and severe (n = 72) COVID-19. We found no significant difference between the genotypic and allelic frequencies of TNF -308G/A in the groups analyzed; however, both the frequencies of the high expression genotype (TT) (CoV: 13.51% vs. CG: 6.30%; p = 0.003) and the *T allele (CoV: 33.56% vs. CG: 24. 81%; p = 0.001) of the IFNG +874T/A polymorphism were higher in the COVID-19 group than in the control group, with no differences between the subgroups of patients with nonsevere and severe COVID-19. The *T allele of IFNG +874T/A (rs2430561) is associated with susceptibility to symptomatic COVID-19. These SNPs provided valuables clues about the potential mechanism involved in the susceptibility to developing symptomatic COVID-19.
Full article
(This article belongs to the Special Issue Basic Sciences for the Conquest of COVID-19)
Open AccessArticle
Characterizing Infections in Two Epidemic Waves of SARS-CoV-2 Omicron Variants: A Cohort Study in Guangzhou, China
by
Lin Qu, Chunyan Xie, Ming Qiu, Lina Yi, Zhe Liu, Lirong Zou, Pei Hu, Huimin Jiang, Huimin Lian, Mingda Yang, Haiyi Yang, Huiling Zeng, Huimin Chen, Jianguo Zhao, Jianpeng Xiao, Jianfeng He, Ying Yang, Liang Chen, Baisheng Li, Jiufeng Sun and Jing Luadd
Show full author list
remove
Hide full author list
Viruses 2024, 16(4), 649; https://doi.org/10.3390/v16040649 - 22 Apr 2024
Abstract
Background: After the adjustment of COVID-19 epidemic policy, mainland China experienced two consecutive waves of Omicron variants within a seven-month period. In Guangzhou city, as one of the most populous regions, the viral infection characteristics, molecular epidemiology, and the dynamic of population immunity
[...] Read more.
Background: After the adjustment of COVID-19 epidemic policy, mainland China experienced two consecutive waves of Omicron variants within a seven-month period. In Guangzhou city, as one of the most populous regions, the viral infection characteristics, molecular epidemiology, and the dynamic of population immunity are still elusive. Methods: We launched a prospective cohort study in the Guangdong Provincial CDC from December 2022 to July 2023. Fifty participants who received the same vaccination regimen and had no previous infection were recruited. Results: 90% of individuals were infected with Omicron BA.5* variants within three weeks in the first wave. Thirteen cases (28.26%) experienced infection with XBB.1* variants, occurring from 14 weeks to 21 weeks after the first wave. BA.5* infections exhibited higher viral loads in nasopharyngeal sites compared to oropharyngeal sites. Compared to BA.5* infections, the XBB.1* infections had significantly milder clinical symptoms, lower viral loads, and shorter durations of virus positivity. The infection with the BA.5* variant elicited varying levels of neutralizing antibodies against XBB.1* among different individuals, even with similar levels of BA.5* antibodies. The level of neutralizing antibodies specific to XBB.1* determined the risk of reinfection. Conclusions: The rapid large-scale infections of the Omicron variants have quickly established herd immunity among the population in mainland China. In the future of the COVID-19 epidemic, a lower infection rate but a longer duration can be expected. Given the large population size and ongoing diversified herd immunity, it remains crucial to closely monitor the molecular epidemiology of SARS-CoV-2 for the emergence of new variants of concern in this region. Additionally, the timely evaluation of the immune status across different age groups is essential for informing future vaccination strategies and intervention policies.
Full article
(This article belongs to the Special Issue COVID-19 Diagnostics in Clinical Applications and Pandemic Controls 2024)
►▼
Show Figures
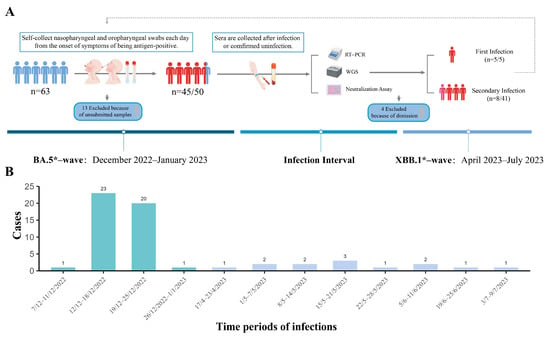
Figure 1
Open AccessArticle
Sosuga Virus Detected in Egyptian Rousette Bats (Rousettus aegyptiacus) in Sierra Leone
by
Brian R. Amman, Alusine H. Koroma, Amy J. Schuh, Immah Conteh, Tara K. Sealy, Ibrahim Foday, Jonathan Johnny, Ibrahim A. Bakarr, Shannon L. M. Whitmer, Emily A. Wright, Aiah A. Gbakima, James Graziano, Camilla Bangura, Emmanuel Kamanda, Augustus Osborne, Emmanuel Saidu, Jonathan A. Musa, Doris F. Bangura, Sammuel M. T. Williams, George M. Fefegula, Christian Sumaila, Juliet Jabaty, Fatmata H. James, Amara Jambai, Kate Garnett, Thomas F. Kamara, Jonathan S. Towner and Aiah Lebbieadd
Show full author list
remove
Hide full author list
Viruses 2024, 16(4), 648; https://doi.org/10.3390/v16040648 - 22 Apr 2024
Abstract
Sosuga virus (SOSV), a rare human pathogenic paramyxovirus, was first discovered in 2012 when a person became ill after working in South Sudan and Uganda. During an ecological investigation, several species of bats were sampled and tested for SOSV RNA and only one
[...] Read more.
Sosuga virus (SOSV), a rare human pathogenic paramyxovirus, was first discovered in 2012 when a person became ill after working in South Sudan and Uganda. During an ecological investigation, several species of bats were sampled and tested for SOSV RNA and only one species, the Egyptian rousette bat (ERBs; Rousettus aegyptiacus), tested positive. Since that time, multiple other species have been sampled and ERBs in Uganda have continued to be the only species of bat positive for SOSV infection. Subsequent studies of ERBs with SOSV demonstrated that ERBs are a competent host for SOSV and shed this infectious virus while exhibiting only minor infection-associated pathology. Following the 2014 Ebola outbreak in West Africa, surveillance efforts focused on discovering reservoirs for zoonotic pathogens resulted in the capture and testing of many bat species. Here, SOSV RNA was detected by qRT-PCR only in ERBs captured in the Moyamba District of Sierra Leone in the central region of the country. These findings represent a substantial range extension from East Africa to West Africa for SOSV, suggesting that this paramyxovirus may occur in ERB populations throughout its sub-Saharan African range.
Full article
(This article belongs to the Special Issue Bat- and Rodent-Borne Zoonotic Viruses)
►▼
Show Figures
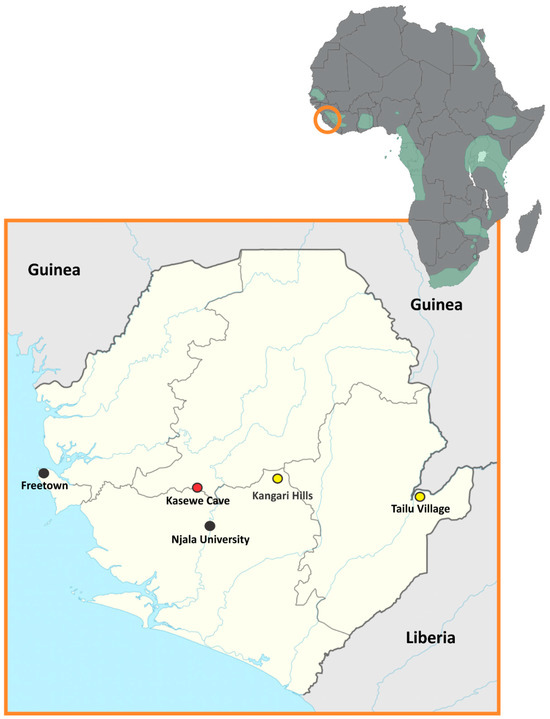
Figure 1
Open AccessReview
Back to the Basics of SARS-CoV-2 Biochemistry: Microvascular Occlusive Glycan Bindings Govern Its Morbidities and Inform Therapeutic Responses
by
David E. Scheim, Peter I. Parry, David J. Rabbolini, Colleen Aldous, Morimasa Yagisawa, Robert Clancy, Thomas J. Borody and Wendy E. Hoy
Viruses 2024, 16(4), 647; https://doi.org/10.3390/v16040647 - 22 Apr 2024
Abstract
Consistent with the biochemistry of coronaviruses as well established over decades, SARS-CoV-2 makes its initial attachment to host cells through the binding of its spike protein (SP) to sialylated glycans (containing the monosaccharide sialic acid) on the cell surface. The virus can then
[...] Read more.
Consistent with the biochemistry of coronaviruses as well established over decades, SARS-CoV-2 makes its initial attachment to host cells through the binding of its spike protein (SP) to sialylated glycans (containing the monosaccharide sialic acid) on the cell surface. The virus can then slide over and enter via ACE2. SARS-CoV-2 SP attaches particularly tightly to the trillions of red blood cells (RBCs), platelets and endothelial cells in the human body, each cell very densely coated with sialic acid surface molecules but having no ACE2 or minimal ACE2. These interlaced attachments trigger the blood cell aggregation, microvascular occlusion and vascular damage that underlie the hypoxia, blood clotting and related morbidities of severe COVID-19. Notably, the two human betacoronaviruses that express a sialic acid-cleaving enzyme are benign, while the other three—SARS, SARS-CoV-2 and MERS—are virulent. RBC aggregation experimentally induced in several animal species using an injected polysaccharide caused most of the same morbidities of severe COVID-19. This glycan biochemistry is key to disentangling controversies that have arisen over the efficacy of certain generic COVID-19 treatment agents and the safety of SP-based COVID-19 vaccines. More broadly, disregard for the active physiological role of RBCs yields unreliable or erroneous reporting of pharmacokinetic parameters as routinely obtained for most drugs and other bioactive agents using detection in plasma, with whole-blood levels being up to 30-fold higher. Appreciation of the active role of RBCs can elucidate the microvascular underpinnings of other health conditions, including cardiovascular disease, and therapeutic opportunities to address them.
Full article
(This article belongs to the Special Issue Glycans in Virus-Host Interactions)
►▼
Show Figures
Figure 1
Open AccessArticle
PCV2 Induced Endothelial Derived IL-8 Affects MoDCs Maturation Mainly via NF-κB Signaling Pathway
by
Mengyu Zhang, Weicheng Xu, Ning Yang, Zhuowei Li, Shuanghai Zhou, Xuewei Liu, Jianfang Wang and Huanrong Li
Viruses 2024, 16(4), 646; https://doi.org/10.3390/v16040646 - 22 Apr 2024
Abstract
Porcine circovirus type 2 (PCV2) infection can cause immunosuppressive diseases in pigs. Vascular endothelial cells (VECs), as the target cells for PCV2, play an important role in the immune response and inflammatory regulation. Endothelial IL-8, which is produced by porcine hip artery endothelial
[...] Read more.
Porcine circovirus type 2 (PCV2) infection can cause immunosuppressive diseases in pigs. Vascular endothelial cells (VECs), as the target cells for PCV2, play an important role in the immune response and inflammatory regulation. Endothelial IL-8, which is produced by porcine hip artery endothelial cells (PIECs) infected with PCV2, can inhibit the maturation of monocyte-derived dendritic cells (MoDCs). Here, we established a co-culture system of MoDCs and different groups of PIECs to further investigate the PCV2-induced endothelial IL-8 signaling pathway that drives the inhibition of MoDC maturation. The differentially expressed genes related to MoDC maturation were mainly enriched in the NF-κB and JAK2-STAT3 signaling pathways. Both the NF-κB related factor RELA and JAK2-STAT3 signaling pathway related factors (IL2RA, JAK, STAT2, STAT5, IL23A, IL7, etc.) decreased significantly in the IL-8 up-regulated group, and increased significantly in the down-regulated group. The expression of NF-κB p65 in the IL-8 up-regulated group was reduced significantly, and the expression of IκBα was increased significantly. Nuclear translocation of NF-κB p65 was inhibited, while the nuclear translocation of p-STAT3 was increased in MoDCs in the PCV2-induced endothelial IL-8 group. The results of treatment with NF-κB signaling pathway inhibitors showed that the maturation of MoDCs was inhibited and the expression of IL-12 and GM-CSF at mRNA level were lower. Inhibition of the JAK2-STAT3 signaling pathway had no significant effect on maturation, and the expression of IL-12 and GM-CSF at mRNA level produced no significant change. In summary, the NF-κB signaling pathway is the main signaling pathway of MoDC maturation, and is inhibited by the PCV2-induced up-regulation of endothelial-derived IL-8.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures
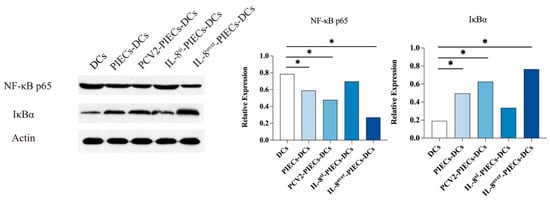
Figure 1
Open AccessArticle
Lysis Physiology of Pseudomonas aeruginosa Infected with ssRNA Phage PRR1
by
Rimantas Daugelavičius, Greta Daujotaitė and Dennis H. Bamford
Viruses 2024, 16(4), 645; https://doi.org/10.3390/v16040645 - 21 Apr 2024
Abstract
The phage PRR1 belongs to the Leviviridae family, a group of ssRNA bacteriophages that infect Gram-negative bacteria. The variety of host cells is determined by the specificity of PRR1 to a pilus encoded by a broad host range of IncP-type plasmids that confer
[...] Read more.
The phage PRR1 belongs to the Leviviridae family, a group of ssRNA bacteriophages that infect Gram-negative bacteria. The variety of host cells is determined by the specificity of PRR1 to a pilus encoded by a broad host range of IncP-type plasmids that confer multiple types of antibiotic resistance to the host. Using P. aeruginosa strain PAO1 as a host, we analyzed the PRR1 infection cycle, focusing on cell lysis. PRR1 infection renders P. aeruginosa cells sensitive to lysozyme approximately 20 min before the start of a drop in suspension turbidity. At the same time, infected cells start to accumulate lipophilic anions. The on-line monitoring of the entire infection cycle showed that single-gene-mediated lysis strongly depends on the host cells’ physiological state. The blockage of respiration or a reduction in the intracellular ATP concentration during the infection resulted in the inhibition of lysis. The same effect was observed when the synthesis of PRR1 lysis protein was induced in an E. coli expression system. In addition, lysis was strongly dependent on the level of aeration. Dissolved oxygen concentrations sufficient to support cell growth did not ensure efficient lysis, and a coupling between cell lysis initiation and aeration level was observed. However, the duration of the drop in suspension turbidity did not depend on the level of aeration.
Full article
(This article belongs to the Special Issue Phage Assembly Pathways — to the Memory of Lindsay Black 2.0)
►▼
Show Figures
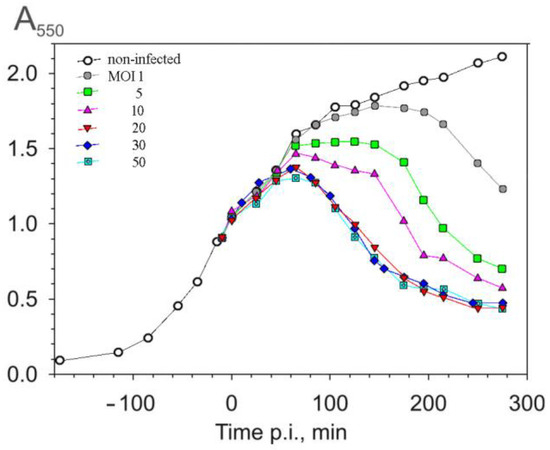
Figure 1
Open AccessArticle
Accumulation Dynamics of Defective Genomes during Experimental Evolution of Two Betacoronaviruses
by
Julia Hillung, María J. Olmo-Uceda, Juan C. Muñoz-Sánchez and Santiago F. Elena
Viruses 2024, 16(4), 644; https://doi.org/10.3390/v16040644 - 20 Apr 2024
Abstract
Virus-encoded replicases often generate aberrant RNA genomes, known as defective viral genomes (DVGs). When co-infected with a helper virus providing necessary proteins, DVGs can multiply and spread. While DVGs depend on the helper virus for propagation, they can in some cases disrupt infectious
[...] Read more.
Virus-encoded replicases often generate aberrant RNA genomes, known as defective viral genomes (DVGs). When co-infected with a helper virus providing necessary proteins, DVGs can multiply and spread. While DVGs depend on the helper virus for propagation, they can in some cases disrupt infectious virus replication, impact immune responses, and affect viral persistence or evolution. Understanding the dynamics of DVGs alongside standard viral genomes during infection remains unclear. To address this, we conducted a long-term experimental evolution of two betacoronaviruses, the human coronavirus OC43 (HCoV-OC43) and the murine hepatitis virus (MHV), in cell culture at both high and low multiplicities of infection (MOI). We then performed RNA-seq at regular time intervals, reconstructed DVGs, and analyzed their accumulation dynamics. Our findings indicate that DVGs evolved to exhibit greater diversity and abundance, with deletions and insertions being the most common types. Notably, some high MOI deletions showed very limited temporary existence, while others became prevalent over time. We observed differences in DVG abundance between high and low MOI conditions in HCoV-OC43 samples. The size distribution of HCoV-OC43 genomes with deletions differed between high and low MOI passages. In low MOI lineages, short and long DVGs were the most common, with an additional cluster in high MOI lineages which became more prevalent along evolutionary time. MHV also showed variations in DVG size distribution at different MOI conditions, though they were less pronounced compared to HCoV-OC43, suggesting a more random distribution of DVG sizes. We identified hotspot regions for deletions that evolved at a high MOI, primarily within cistrons encoding structural and accessory proteins. In conclusion, our study illustrates the widespread formation of DVGs during betacoronavirus evolution, influenced by MOI and cell- and virus-specific factors.
Full article
(This article belongs to the Special Issue Viruses 2024 - A World of Viruses)
Open AccessArticle
Autophagy Deregulation in HIV-1-Infected Cells Increases Extracellular Vesicle Release and Contributes to TLR3 Activation
by
Catherine DeMarino, Maria Cowen, Anastasia Williams, Pooja Khatkar, Fardokht A. Abulwerdi, Lisa Henderson, Julia Denniss, Michelle L. Pleet, Delores R. Luttrell, Iosif Vaisman, Lance A. Liotta, Joseph Steiner, Stuart F. J. Le Grice, Avindra Nath and Fatah Kashanchi
Viruses 2024, 16(4), 643; https://doi.org/10.3390/v16040643 - 20 Apr 2024
Abstract
Human immunodeficiency virus type 1 (HIV-1) infection can result in HIV-associated neurocognitive disorder (HAND), a spectrum of disorders characterized by neurological impairment and chronic inflammation. Combined antiretroviral therapy (cART) has elicited a marked reduction in the number of individuals diagnosed with HAND. However,
[...] Read more.
Human immunodeficiency virus type 1 (HIV-1) infection can result in HIV-associated neurocognitive disorder (HAND), a spectrum of disorders characterized by neurological impairment and chronic inflammation. Combined antiretroviral therapy (cART) has elicited a marked reduction in the number of individuals diagnosed with HAND. However, there is continual, low-level viral transcription due to the lack of a transcription inhibitor in cART regimens, which results in the accumulation of viral products within infected cells. To alleviate stress, infected cells can release accumulated products, such as TAR RNA, in extracellular vesicles (EVs), which can contribute to pathogenesis in neighboring cells. Here, we demonstrate that cART can contribute to autophagy deregulation in infected cells and increased EV release. The impact of EVs released from HIV-1 infected myeloid cells was found to contribute to CNS pathogenesis, potentially through EV-mediated TLR3 (Toll-like receptor 3) activation, suggesting the need for therapeutics to target this mechanism. Three HIV-1 TAR-binding compounds, 103FA, 111FA, and Ral HCl, were identified that recognize TAR RNA and reduce TLR activation. These data indicate that packaging of viral products into EVs, potentially exacerbated by antiretroviral therapeutics, may induce chronic inflammation of the CNS observed in cART-treated patients, and novel therapeutic strategies may be exploited to mitigate morbidity.
Full article
(This article belongs to the Special Issue HIV-1 Infection and Immunometabolism: Relevance to HIV Pathogenesis and Persistence 2.0)
►▼
Show Figures
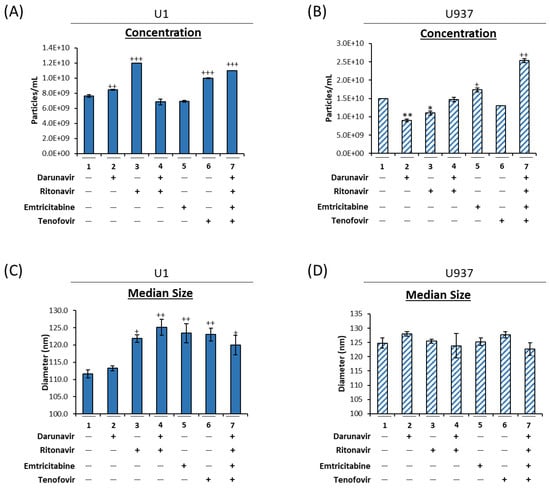
Figure 1
Open AccessArticle
Monitoring HPV Prevalence and Risk Cofactors for Abnormal Cytology in the Post-Vaccination Period among Croatian Women
by
Ena Pešut, Ivana Šimić, Rajko Fureš, Nina Milutin Gašperov, Cvjetko Lež, Fabijan Feratović, Tomica Kukina Žvigač, Magdalena Grce, Ivana Erceg Ivkošić and Ivan Sabol
Viruses 2024, 16(4), 642; https://doi.org/10.3390/v16040642 - 20 Apr 2024
Abstract
The incidence and mortality rate of cervical cancer in Croatia remains a health challenge despite screening efforts. Besides the persistent infection with HPV, the development of cancer is also associated with some cofactors. The goal of this study was to assess circulating HPV
[...] Read more.
The incidence and mortality rate of cervical cancer in Croatia remains a health challenge despite screening efforts. Besides the persistent infection with HPV, the development of cancer is also associated with some cofactors. The goal of this study was to assess circulating HPV genotypes and risk factors for the development of cervical precancer after almost 16 years from the onset of HPV vaccination in Croatia. In this study, a total of 321 women attending gynecological care were evaluated. Relevant medical and demographic information, including cytology, were collected. HPV genotyping was performed by PCR. Comparing the HPV types found in circulation in the pre-vaccination (1999–2015) and post-vaccination periods (2020–2023), a statistically significant reduction in HPV 31 was noted, while the overall prevalence increased in the post-vaccination period. Besides the expected HPV positivity as a risk factor, the history of smoking was associated with LSIL or worse cytology at enrollment. For the first time, this population study revealed a statistically significant shift in the HPV genotype in the post-vaccination period, as well as the confirmation of risk factors for the development of abnormal cytology among Croatian women.
Full article
(This article belongs to the Special Issue Human and Animal Papillomavirus: Infections, Genetics, and Vaccines)
►▼
Show Figures

Figure 1
Open AccessArticle
New Genera and Species of Caulobacter and Brevundimonas Bacteriophages Provide Insights into Phage Genome Evolution
by
Bert Ely, Michael Hils, Aaron Clarke, Maegan Albert, Nadia Holness, Jacob Lenski and Tannaz Mohammadi
Viruses 2024, 16(4), 641; https://doi.org/10.3390/v16040641 - 20 Apr 2024
Abstract
Previous studies have identified diverse bacteriophages that infect Caulobacter vibrioides strain CB15 ranging from small RNA phages to four genera of jumbo phages. In this study, we focus on 20 bacteriophages whose genomes range from 40 to 60 kb in length. Genome comparisons
[...] Read more.
Previous studies have identified diverse bacteriophages that infect Caulobacter vibrioides strain CB15 ranging from small RNA phages to four genera of jumbo phages. In this study, we focus on 20 bacteriophages whose genomes range from 40 to 60 kb in length. Genome comparisons indicated that these diverse phages represent six Caulobacter phage genera and one additional genus that includes both Caulobacter and Brevundimonas phages. Within species, comparisons revealed that both single base changes and inserted or deleted genetic material cause the genomes of closely related phages to diverge. Among genera, the basic gene order and the orientation of key genes were retained with most of the observed variation occurring at ends of the genomes. We hypothesize that the nucleotide sequences of the ends of these phage genomes are less important than the need to maintain the size of the genome and the stability of the corresponding mRNAs.
Full article
(This article belongs to the Special Issue Bacteriophage Diversity)
►▼
Show Figures

Figure 1
Open AccessArticle
The Dual-Targeted Fusion Inhibitor Clofazimine Binds to the S2 Segment of the SARS-CoV-2 Spike Protein
by
Matthew R. Freidel, Pratiti A. Vakhariya, Shalinder K. Sardarni and Roger S. Armen
Viruses 2024, 16(4), 640; https://doi.org/10.3390/v16040640 - 20 Apr 2024
Abstract
Clofazimine and Arbidol have both been reported to be effective in vitro SARS-CoV-2 fusion inhibitors. Both are promising drugs that have been repurposed for the treatment of COVID-19 and have been used in several previous and ongoing clinical trials. Small-molecule bindings to expressed
[...] Read more.
Clofazimine and Arbidol have both been reported to be effective in vitro SARS-CoV-2 fusion inhibitors. Both are promising drugs that have been repurposed for the treatment of COVID-19 and have been used in several previous and ongoing clinical trials. Small-molecule bindings to expressed constructs of the trimeric S2 segment of Spike and the full-length SARS-CoV-2 Spike protein were measured using a Surface Plasmon Resonance (SPR) binding assay. We demonstrate that Clofazimine, Toremifene, Arbidol and its derivatives bind to the S2 segment of the Spike protein. Clofazimine provided the most reliable and highest-quality SPR data for binding with S2 over the conditions explored. A molecular docking approach was used to identify the most favorable binding sites on the S2 segment in the prefusion conformation, highlighting two possible small-molecule binding sites for fusion inhibitors. Results related to molecular docking and modeling of the structure–activity relationship (SAR) of a newly reported series of Clofazimine derivatives support the proposed Clofazimine binding site on the S2 segment. When the proposed Clofazimine binding site is superimposed with other experimentally determined coronavirus structures in structure–sequence alignments, the changes in sequence and structure may rationalize the broad-spectrum antiviral activity of Clofazimine in closely related coronaviruses such as SARS-CoV, MERS, hCoV-229E, and hCoV-OC43.
Full article
(This article belongs to the Special Issue Innovative Drug Discovery for Emerging Viral Diseases)
►▼
Show Figures
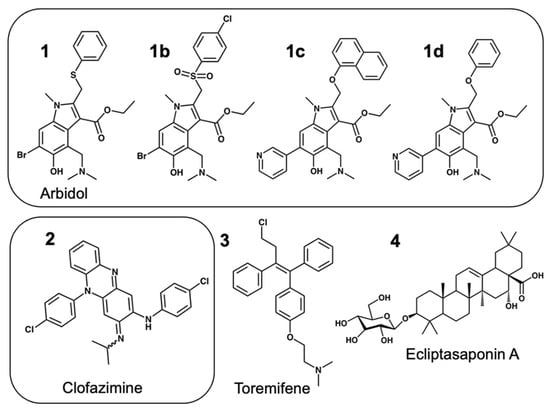
Figure 1
Open AccessArticle
Vitamin D Levels and SARS-CoV-2 Infection among Medically Underserved Populations in the Minority and Rural Coronavirus Insights Study
by
Makella S. Coudray, Shantoy Hansel, Salvatore Alesci, William A. Meyer III, Robert H. Christenson, Latrice G. Landry, Christina Edwards, Gary Puckrein, Derrick J. Forney and Ola Akinboboye
Viruses 2024, 16(4), 639; https://doi.org/10.3390/v16040639 - 19 Apr 2024
Abstract
Background: Extant literature presents contradictory findings on the role of vitamin D on SARS-CoV-2 infection. Our study included an examination of the relationship between vitamin D levels and SARS-CoV-2 infection among the Minority and Rural Coronavirus Insights Study (MRCIS) cohort, a diverse population
[...] Read more.
Background: Extant literature presents contradictory findings on the role of vitamin D on SARS-CoV-2 infection. Our study included an examination of the relationship between vitamin D levels and SARS-CoV-2 infection among the Minority and Rural Coronavirus Insights Study (MRCIS) cohort, a diverse population of medically underserved persons presenting at five Federally qualified health centers in the United States. Methods: We conducted a descriptive analysis to explore the relationship between vitamin D levels and SARS-CoV-2 infection among medically underserved participants. A combined molecular and serologic assessment was used to determine the prevalence of SARS-CoV-2 infection. Vitamin D was examined as both a categorical (vitamin D status: deficient, insufficient, optimal) and continuous (vitamin D level) variable. Chi-squared testing, polynomial regression models, and logistic regression models were used to assess the relationship between vitamin D and SARS-CoV-2 infection. Results: The overall SARS-CoV-2 infection rate among participants was 25.9%. Most participants were either vitamin D deficient (46.5%) or insufficient (29.7%), and 23.8% had an optimal level. Vitamin D status was significantly associated with key SARS-CoV-2 infection risk factors. As mean vitamin D levels increased, the proportion of participants with SARS-CoV-2 infection decreased. For every 10 ng/mL increase in vitamin D levels the odds of SARS-CoV-2 infection decreased by 12% when adjusting for race/ethnicity and age (main effect model). Participants who identified as Hispanic/Latino or Black non-Hispanic had approximately two times increased odds of SARS-CoV-2 infection when adjusting for age and vitamin D levels compared to white non-Hispanics. However, when additional factors were added to the main effect model, the relationship between vitamin D levels and SARS-CoV-2 infection did not remain significant. Conclusion: Vitamin D levels were associated with an increased risk of SARS-CoV-2 infection. Hispanic/Latino and Black, non-Hispanic compared to White, non-Hispanic participants were at increased odds for infection, after adjusting for race/ethnicity and age.
Full article
(This article belongs to the Special Issue Opportunistic Viral Infections 2nd Edition)
►▼
Show Figures
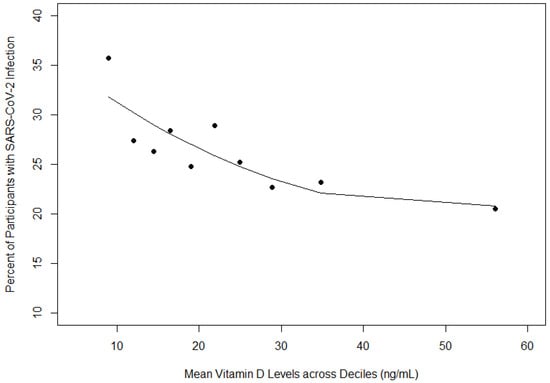
Figure 1

Journal Menu
► ▼ Journal Menu-
- Viruses Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Diseases, Infectious Disease Reports, Pathogens, Viruses, TropicalMed
Human Monkeypox Research
Topic Editors: Shailendra K. Saxena, Ahmed Sayed Abdel-MoneimDeadline: 30 June 2024
Topic in
Biomedicines, JCM, Pathogens, Vaccines, Viruses
Discovery and Development of Monkeypox Disease Treatments
Topic Editors: Mohd Imran, Ali A. RabaanDeadline: 31 August 2024
Topic in
Brain Sciences, Clinics and Practice, COVID, Life, Vaccines, Viruses
Multifaceted Efforts from Basic Research to Clinical Practice in Controlling COVID-19 Disease
Topic Editors: Yih-Horng Shiao, Rashi OjhaDeadline: 30 September 2024

Conferences
Special Issues
Special Issue in
Viruses
Oncolytic Viruses as Immunotherapeutic Agents
Guest Editor: Nadine Van MontfoortDeadline: 1 May 2024
Special Issue in
Viruses
Animal Coronaviruses: Infection, Prevention, and Antivirals
Guest Editor: Tomomi TakanoDeadline: 20 May 2024
Special Issue in
Viruses
Bacteriophages and Biofilms 2.0
Guest Editors: Zuzanna Drulis-Kawa, Tomasz OlszakDeadline: 31 May 2024
Special Issue in
Viruses
The Inflammasomes - Key Players in Antiviral Response
Guest Editors: Dong-Yan Jin, Tsan Sam XiaoDeadline: 15 June 2024
Topical Collections
Topical Collection in
Viruses
Poxviruses
Collection Editors: Giliane de Souza Trindade, Galileu Barbosa Costa, Flavio Guimaraes da Fonseca
Topical Collection in
Viruses
Coronaviruses
Collection Editors: Luis Martinez-Sobrido, Fernando Almazan Toral
Topical Collection in
Viruses
SARS-CoV-2 and COVID-19
Collection Editors: Luis Martinez-Sobrido, Fernando Almazan Toral








