Journal Description
Genes
Genes
is a peer-reviewed, open access journal of genetics and genomics published monthly online by MDPI. The Spanish Society for Biochemistry and Molecular Biology (SEBBM) is affiliated with Genes and their members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, and other databases.
- Journal Rank: JCR - Q2 (Genetics & Heredity) / CiteScore - Q2 (Genetics)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.5 days after submission; acceptance to publication is undertaken in 2.3 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: Reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
3.5 (2022);
5-Year Impact Factor:
3.9 (2022)
Latest Articles
WGCNA Reveals Hub Genes and Key Gene Regulatory Pathways of the Response of Soybean to Infection by Soybean mosaic virus
Genes 2024, 15(5), 566; https://doi.org/10.3390/genes15050566 (registering DOI) - 27 Apr 2024
Abstract
Soybean mosaic virus (SMV) is one of the main pathogens that can negatively affect soybean production and quality. To study the gene regulatory network of soybeans in response to SMV SC15, the resistant line X149 and susceptible line X97 were subjected to transcriptome
[...] Read more.
Soybean mosaic virus (SMV) is one of the main pathogens that can negatively affect soybean production and quality. To study the gene regulatory network of soybeans in response to SMV SC15, the resistant line X149 and susceptible line X97 were subjected to transcriptome analysis at 0, 2, 8, 12, 24, and 48 h post-inoculation (hpi). Differential expression analysis revealed that 10,190 differentially expressed genes (DEGs) responded to SC15 infection. Weighted gene co-expression network analysis (WGCNA) was performed to identify highly related resistance gene modules; in total, eight modules, including 2256 DEGs, were identified. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of 2256 DEGs revealed that the genes significantly clustered into resistance-related pathways, such as the plant–pathogen interaction pathway, mitogen-activated protein kinases (MAPK) signaling pathway, and plant hormone signal transduction pathway. Among these pathways, we found that the flg22, Ca2+, hydrogen peroxide (H2O2), and abscisic acid (ABA) regulatory pathways were fully covered by 36 DEGs. Among the 36 DEGs, the gene Glyma.01G225100 (protein phosphatase 2C, PP2C) in the ABA regulatory pathway, the gene Glyma.16G031900 (WRKY transcription factor 22, WRKY22) in Ca2+ and H2O2 regulatory pathways, and the gene Glyma.04G175300 (calcium-dependent protein kinase, CDPK) in Ca2+ regulatory pathways were highly connected hub genes. These results indicate that the resistance of X149 to SC15 may depend on the positive regulation of flg22, Ca2+, H2O2, and ABA regulatory pathways. Our study further showed that superoxide dismutase (SOD) activity, H2O2 content, and catalase (CAT) and peroxidase (POD) activities were significantly up-regulated in the resistant line X149 compared with those in 0 hpi. This finding indicates that the H2O2 regulatory pathway might be dependent on flg22- and Ca2+-pathway-induced ROS generation. In addition, two hub genes, Glyma.07G190100 (encoding F-box protein) and Glyma.12G185400 (encoding calmodulin-like proteins, CMLs), were also identified and they could positively regulate X149 resistance. This study provides pathways for further investigation of SMV resistance mechanisms in soybean.
Full article
(This article belongs to the Section Plant Genetics and Genomics)
Open AccessBrief Report
Examining the Effect of Genes on Depression as Mediated by Smoking and Modified by Sex
by
Kirsten Voorhies, Julian Hecker, Sanghun Lee, Georg Hahn, Dmitry Prokopenko, Merry-Lynn McDonald, Alexander C. Wu, Ann Wu, John E. Hokanson, Michael H. Cho, Christoph Lange, Karin F. Hoth and Sharon M. Lutz
Genes 2024, 15(5), 565; https://doi.org/10.3390/genes15050565 (registering DOI) - 27 Apr 2024
Abstract
Depression is heritable, differs by sex, and has environmental risk factors such as cigarette smoking. However, the effect of single nucleotide polymorphisms (SNPs) on depression through cigarette smoking and the role of sex is unclear. In order to examine the association of SNPs
[...] Read more.
Depression is heritable, differs by sex, and has environmental risk factors such as cigarette smoking. However, the effect of single nucleotide polymorphisms (SNPs) on depression through cigarette smoking and the role of sex is unclear. In order to examine the association of SNPs with depression and smoking in the UK Biobank with replication in the COPDGene study, we used counterfactual-based mediation analysis to test the indirect or mediated effect of SNPs on broad depression through the log of pack-years of cigarette smoking, adjusting for age, sex, current smoking status, and genetic ancestry (via principal components). In secondary analyses, we adjusted for age, sex, current smoking status, genetic ancestry (via principal components), income, education, and living status (urban vs. rural). In addition, we examined sex-stratified mediation models and sex-moderated mediation models. For both analyses, we adjusted for age, current smoking status, and genetic ancestry (via principal components). In the UK Biobank, rs6424532 [LOC105378800] had a statistically significant indirect effect on broad depression through the log of pack-years of cigarette smoking (p = 4.0 × 10−4) among all participants and a marginally significant indirect effect among females (p = 0.02) and males (p = 4.0 × 10−3). Moreover, rs10501696 [GRM5] had a marginally significant indirect effect on broad depression through the log of pack-years of cigarette smoking (p = 0.01) among all participants and a significant indirect effect among females (p = 2.2 × 10−3). In the secondary analyses, the sex-moderated indirect effect was marginally significant for rs10501696 [GRM5] on broad depression through the log of pack-years of cigarette smoking (p = 0.01). In the COPDGene study, the effect of an SNP (rs10501696) in GRM5 on depressive symptoms and medication was mediated by log of pack-years (p = 0.02); however, no SNPs had a sex-moderated mediated effect on depressive symptoms. In the UK Biobank, we found SNPs in two genes [LOC105378800, GRM5] with an indirect effect on broad depression through the log of pack-years of cigarette smoking. In addition, the indirect effect for GRM5 on broad depression through smoking may be moderated by sex. These results suggest that genetic regions associated with broad depression may be mediated by cigarette smoking and this relationship may be moderated by sex.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
Open AccessReview
Progress in Rice Breeding Based on Genomic Research
by
Xingye Yang, Shicong Yu, Shen Yan, Hao Wang, Wei Fang, Yanqing Chen, Xiaoding Ma and Longzhi Han
Genes 2024, 15(5), 564; https://doi.org/10.3390/genes15050564 (registering DOI) - 27 Apr 2024
Abstract
The role of rice genomics in breeding progress is becoming increasingly important. Deeper research into the rice genome will contribute to the identification and utilization of outstanding functional genes, enriching the diversity and genetic basis of breeding materials and meeting the diverse demands
[...] Read more.
The role of rice genomics in breeding progress is becoming increasingly important. Deeper research into the rice genome will contribute to the identification and utilization of outstanding functional genes, enriching the diversity and genetic basis of breeding materials and meeting the diverse demands for various improvements. Here, we review the significant contributions of rice genomics research to breeding progress over the last 25 years, discussing the profound impact of genomics on rice-genome sequencing, functional-gene exploration, and novel breeding methods, and we provide valuable insights for future research and breeding practices.
Full article
(This article belongs to the Special Issue Genomic Studies of Plant Breeding)
Open AccessReview
Roles of Nuclear Orphan Receptors TR2 and TR4 during Hematopoiesis
by
Greggory Myers, Yanan Sun, Yu Wang, Hajar Benmhammed and Shuaiying Cui
Genes 2024, 15(5), 563; https://doi.org/10.3390/genes15050563 (registering DOI) - 27 Apr 2024
Abstract
TR2 and TR4 (NR2C1 and NR2C2, respectively) are evolutionarily conserved nuclear orphan receptors capable of binding direct repeat sequences in a stage-specific manner. Like other nuclear receptors, TR2 and TR4 possess important roles in transcriptional activation or repression with developmental stage and tissue
[...] Read more.
TR2 and TR4 (NR2C1 and NR2C2, respectively) are evolutionarily conserved nuclear orphan receptors capable of binding direct repeat sequences in a stage-specific manner. Like other nuclear receptors, TR2 and TR4 possess important roles in transcriptional activation or repression with developmental stage and tissue specificity. TR2 and TR4 bind DNA and possess the ability to complex with available cofactors mediating developmental stage-specific actions in primitive and definitive erythrocytes. In erythropoiesis, TR2 and TR4 are required for erythroid development, maturation, and key erythroid transcription factor regulation. TR2 and TR4 recruit and interact with transcriptional corepressors or coactivators to elicit developmental stage-specific gene regulation during hematopoiesis.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
Open AccessArticle
Decoding Evolution of Rubioideae: Plastomes Reveal Sweet Secrets of Codon Usage, Diagnostides, and Superbarcoding
by
Kamil Ciborowski, Monika Szczecińska, Mateusz Maździarz, Jakub Sawicki and Łukasz Paukszto
Genes 2024, 15(5), 562; https://doi.org/10.3390/genes15050562 (registering DOI) - 27 Apr 2024
Abstract
Galium genus belongs to the Rubiaceae family, which consists of approximately 14,000 species. In comparison to its well-known relatives, the plastomes of the Galium genus have not been explored so far. The plastomes of this genus have a typical, quadripartite structure, but differ
[...] Read more.
Galium genus belongs to the Rubiaceae family, which consists of approximately 14,000 species. In comparison to its well-known relatives, the plastomes of the Galium genus have not been explored so far. The plastomes of this genus have a typical, quadripartite structure, but differ in gene content, since the infA gene is missing in Galium palustre and Galium trfidum. An evaluation of the effectiveness of using entire chloroplast genome sequences as superbarcodes for accurate plant species identification revealed the high potential of this method for molecular delimitation within the genus and tribe. The trnE-UUC—psbD region showed the biggest number of diagnostides (diagnostic nucleotides) which might be new potential barcodes, not only in Galium, but also in other closely related genera. Relative synonymous codon usage (RSCU) appeared to be connected with the phylogeny of the Rubiaceae family, showing that during evolution, plants started preferring specific codons over others.
Full article
(This article belongs to the Special Issue The Evolutionary Genetics and Genomics of Speciation)
Open AccessReview
Sarcopenia as a Risk Factor for Alzheimer’s Disease: Genetic and Epigenetic Perspectives
by
Stuart M. Raleigh and Kayleigh J. A. Orchard
Genes 2024, 15(5), 561; https://doi.org/10.3390/genes15050561 (registering DOI) - 27 Apr 2024
Abstract
Sarcopenia, defined as the age-associated loss of muscle mass and increased fragility with age, is increasing worldwide. The condition often precedes the development of Alzheimer’s disease, thereby decreasing the levels of mobility and physical activity in those affected. Indeed, the loss of muscle
[...] Read more.
Sarcopenia, defined as the age-associated loss of muscle mass and increased fragility with age, is increasing worldwide. The condition often precedes the development of Alzheimer’s disease, thereby decreasing the levels of mobility and physical activity in those affected. Indeed, the loss of muscle mass has, in some studies, been associated with an increased risk of Alzheimer’s disease and other dementias. However, a detailed understanding of the interplay between both conditions is not available and needs to be thoroughly addressed. In the following review, we focus on several genes, specifically APOE, BDNF, ACE, FTO, and FNDC5, that have been associated with both conditions. We also discuss the epigenetic regulation of each of these genes along with non-coding RNAs (ncRNAs) that may have a role in the development of both the sarcopenic and Alzheimer’s disease phenotypes. Finally, we assert that the application of systems biology will unravel the relationship between sarcopenia and Alzheimer’s disease and believe that the prevention of muscle loss in older age will reduce the incidence of debilitating cognitive decline.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
►▼
Show Figures
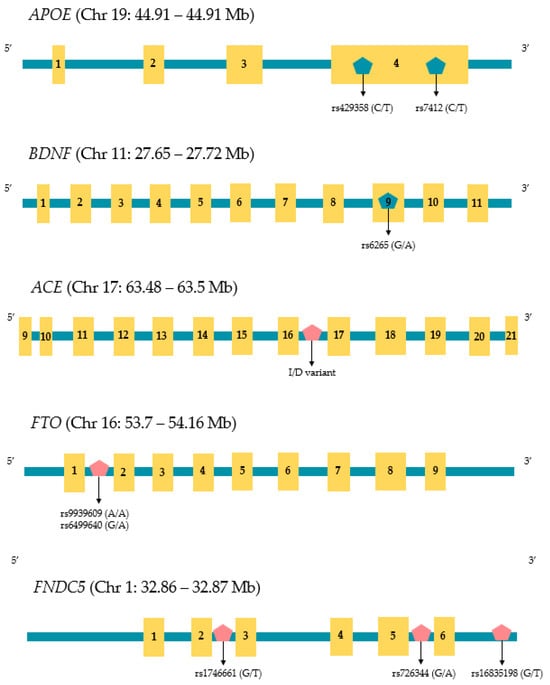
Figure 1
Open AccessArticle
Exploring the Role of E6 and E7 Oncoproteins in Cervical Oncogenesis through MBD2/3-NuRD Complex Chromatin Remodeling
by
Alina Fudulu, Carmen Cristina Diaconu, Iulia Virginia Iancu, Adriana Plesa, Adrian Albulescu, Marinela Bostan, Demetra Gabriela Socolov, Irina Liviana Stoian, Raluca Balan, Gabriela Anton and Anca Botezatu
Genes 2024, 15(5), 560; https://doi.org/10.3390/genes15050560 (registering DOI) - 27 Apr 2024
Abstract
Background: Cervical cancer is among the highest-ranking types of cancer worldwide, with human papillomavirus (HPV) as the agent driving the malignant process. One aspect of the infection’s evolution is given by epigenetic modifications, mainly DNA methylation and chromatin alteration. These processes are guided
[...] Read more.
Background: Cervical cancer is among the highest-ranking types of cancer worldwide, with human papillomavirus (HPV) as the agent driving the malignant process. One aspect of the infection’s evolution is given by epigenetic modifications, mainly DNA methylation and chromatin alteration. These processes are guided by several chromatin remodeling complexes, including NuRD. The purpose of this study was to evaluate the genome-wide binding patterns of the NuRD complex components (MBD2 and MBD3) in the presence of active HPV16 E6 and E7 oncogenes and to determine the potential of identified genes through an experimental model to differentiate between cervical precursor lesions, with the aim of establishing their utility as biomarkers. Methods: The experimental model was built using the CaSki cell line and shRNA for E6 and E7 HPV16 silencing, ChIP-seq, qRT-PCR, and Western blot analyses. Selected genes’ expression was also assessed in patients. Results: Several genes have been identified to exhibit altered transcriptional activity due to the influence of HPV16 E6/E7 viral oncogenes acting through the MBD2/MBD3 NuRD complex, linking them to viral infection and cervical oncogenesis. Conclusions: The impacted genes primarily play roles in governing gene transcription, mRNA processing, and regulation of translation. Understanding these mechanisms offers valuable insights into the process of HPV-induced oncogenesis.
Full article
(This article belongs to the Topic Advances in Genetics and Precision Medicine in Human Diseases)
►▼
Show Figures
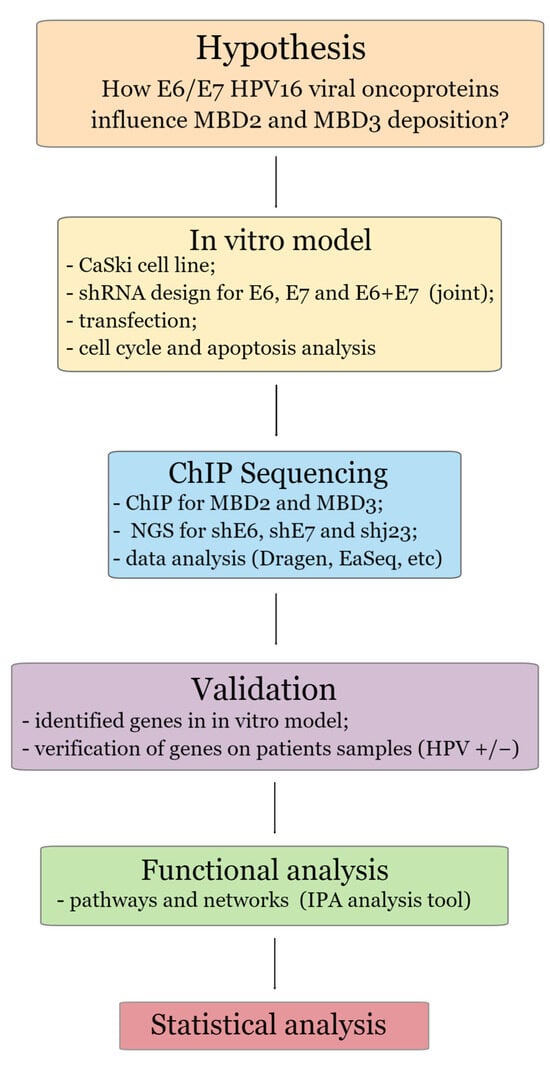
Figure 1
Open AccessArticle
Optical Genome Mapping as a New Tool to Overcome Conventional Cytogenetics Limitations in Patients with Bone Marrow Failure
by
June Iriondo, Ana Gómez, Josune Zubicaray, Jorge Garcia-Martinez, Lorea Abad, Carmen Matesanz, Reyes Giménez, Almudena Galán, Alejandro Sanz, Elena Sebastián, Jesús Gonzalez de Pablo, Ana de la Cruz, Manuel Ramírez and Julián Sevilla
Genes 2024, 15(5), 559; https://doi.org/10.3390/genes15050559 (registering DOI) - 27 Apr 2024
Abstract
Cytogenetic studies are essential in the diagnosis and follow up of patients with bone marrow failure syndromes (BMFSs), but obtaining good quality results is often challenging due to hypocellularity. Optical Genome Mapping (OGM), a novel technology capable of detecting most types chromosomal structural
[...] Read more.
Cytogenetic studies are essential in the diagnosis and follow up of patients with bone marrow failure syndromes (BMFSs), but obtaining good quality results is often challenging due to hypocellularity. Optical Genome Mapping (OGM), a novel technology capable of detecting most types chromosomal structural variants (SVs) at high resolution, is being increasingly used in many settings, including hematologic malignancies. Herein, we compared conventional cytogenetic techniques to OGM in 20 patients with diverse BMFSs. Twenty metaphases for the karyotype were only obtained in three subjects (15%), and no SVs were found in any of the samples. One patient with culture failure showed a gain in chromosome 1q by fluorescence in situ hybridization, which was confirmed by OGM. In contrast, OGM provided good quality results in all subjects, and SVs were detected in 14 of them (70%), mostly corresponding to cryptic submicroscopic alterations not observed by standard techniques. Therefore, OGM emerges as a powerful tool that provides complete and evaluable results in hypocellular BMFSs, reducing multiple tests into a single assay and overcoming some of the main limitations of conventional techniques. Furthermore, in addition to confirming the abnormalities detected by conventional techniques, OGM found new alterations beyond their detection limits.
Full article
(This article belongs to the Special Issue Diagnosis and Therapies for Rare Diseases)
Open AccessReview
Genetic Mechanisms Driving Uterine Leiomyoma Pathobiology, Epidemiology, and Treatment
by
Malini S. Ramaiyer, Eslam Saad, Irem Kurt and Mostafa A. Borahay
Genes 2024, 15(5), 558; https://doi.org/10.3390/genes15050558 (registering DOI) - 27 Apr 2024
Abstract
Uterine leiomyomas (ULs) are the most common benign tumor of the uterus. They can be associated with symptoms including abnormal uterine bleeding, pelvic pain, urinary frequency, and pregnancy complications. Despite the high prevalence of UL, its underlying pathophysiology mechanisms have historically been poorly
[...] Read more.
Uterine leiomyomas (ULs) are the most common benign tumor of the uterus. They can be associated with symptoms including abnormal uterine bleeding, pelvic pain, urinary frequency, and pregnancy complications. Despite the high prevalence of UL, its underlying pathophysiology mechanisms have historically been poorly understood. Several mechanisms of pathogenesis have been suggested, implicating various genes, growth factors, cytokines, chemokines, and microRNA aberrations. The purpose of this study is to summarize the current research on the relationship of genetics with UL. Specifically, we performed a literature review of published studies to identify how genetic aberrations drive pathophysiology, epidemiology, and therapeutic approaches of UL. With regards to pathophysiology, research has identified MED12 mutations, HMGA2 overexpression, fumarate hydratase deficiency, and cytogenetic abnormalities as contributors to the development of UL. Additionally, epigenetic modifications, such as histone acetylation and DNA methylation, have been identified as contributing to UL tumorigenesis. Specifically, UL stem cells have been found to contain a unique DNA methylation pattern compared to more differentiated UL cells, suggesting that DNA methylation has a role in tumorigenesis. On a population level, genome-wide association studies (GWASs) and epidemiologic analyses have identified 23 genetic loci associated with younger age at menarche and UL growth. Additionally, various GWASs have investigated genetic loci as potential drivers of racial disparities in UL incidence. For example, decreased expression of Cytohesin 4 in African Americans has been associated with increased UL risk. Recent studies have investigated various therapeutic options, including ten-eleven translocation proteins mediating DNA methylation, adenovirus vectors for drug delivery, and “suicide gene therapy” to induce apoptosis. Overall, improved understanding of the genetic and epigenetic drivers of UL on an individual and population level can propel the discovery of novel therapeutic options.
Full article
(This article belongs to the Special Issue Genetics and Genomics of Female Reproduction)
►▼
Show Figures

Figure 1
Open AccessArticle
Complete Mitochondrial Genome of Four Peristediidae Fish Species: Genome Characterization and Phylogenetic Analysis
by
Xianhui Liao, Yijia Shih, Chenghao Jia and Tianxiang Gao
Genes 2024, 15(5), 557; https://doi.org/10.3390/genes15050557 (registering DOI) - 27 Apr 2024
Abstract
The systematic revision of the family Peristediidae remains an unresolved issue due to their diverse and unique morphology. Despite the popularity of using mitochondrial genome research to comprehensively understand phylogenetic relationships in fish, genetic data for peristediid fish need to be included. Therefore,
[...] Read more.
The systematic revision of the family Peristediidae remains an unresolved issue due to their diverse and unique morphology. Despite the popularity of using mitochondrial genome research to comprehensively understand phylogenetic relationships in fish, genetic data for peristediid fish need to be included. Therefore, this study aims to investigate the mitochondrial genomic characteristics and intra-family phylogenetic relationships of Peristediidae by utilizing mitochondrial genome analysis. Therefore, this study aims to investigate the phylogenetic relationship of Peristediidae by utilizing mitochondrial genome analysis. The mitochondrial genome of four species of Peristediidae (Peristedion liorhynchus, Satyrichthys welchi, Satyrichthys rieffeli, and Scalicus amiscus) collected in the East China Sea was studied. The mitochondrial gene sequence lengths of four fish species were 16,533 bp, 16,526 bp, 16,527 bp, and 16,526 bp, respectively. They had the same mitochondrial structure and were all composed of 37 genes and one control region. Most PCGs used ATG as the start codon, and a few used GTG as the start codon. An incomplete stop codon (TA/T) occurred. The AT-skew and GC-skew values of 13 PCGs from four species were negative, and the GC-skew amplitude was greater than that of AT-skew. All cases of D-arm were found in tRNA-Ser (GCT). The Ka/Ks ratio analysis indicated that 13 PCGs were suffering purifying selection. Based on 12 PCGs (excluding ND6) sequences, a phylogenetic tree was constructed using Bayesian inference (BI) and maximum likelihood (ML) methods, providing a further supplement to the scientific classification of Peristediidae fish. According to the results of divergence time, the four species of fish had apparent divergence in the Early Cenozoic, which indicates that the geological events at that time caused the climax of species divergence and evolution.
Full article
(This article belongs to the Special Issue DNA Taxonomy, Molecular Phylogeny and Population Genetics of Cartilaginous Fishes and Teleost Fishes)
►▼
Show Figures

Figure 1
Open AccessArticle
The Relationship between miR-5682 and Nutritional Status of Radiotherapy-Treated Male Laryngeal Cancer Patients
by
Marcin Mazurek, Anna Brzozowska, Mirosław Maziarz, Teresa Małecka-Massalska and Tomasz Powrózek
Genes 2024, 15(5), 556; https://doi.org/10.3390/genes15050556 (registering DOI) - 27 Apr 2024
Abstract
Background: Nutritional deficiencies are frequently observed in patients with head and neck cancer (HNC) undergoing radiation therapy. microRNAs (miRNAs) were found to play an important role in the development of metabolic disorders throughout regulation of genes involved in inflammatory responses. This study aimed
[...] Read more.
Background: Nutritional deficiencies are frequently observed in patients with head and neck cancer (HNC) undergoing radiation therapy. microRNAs (miRNAs) were found to play an important role in the development of metabolic disorders throughout regulation of genes involved in inflammatory responses. This study aimed to explore the correlation between pre-treatment miR-5682 expression and parameters reflecting nutritional deficits in laryngeal cancer (LC) patients subjected to radiotherapy (RT). Methods: Expression of miR-5682 was analyzed in plasma samples of 56 male LC individuals. Nutritional status of LC patients was assessed using anthropometric and laboratory parameters, bioelectrical impedance analysis (BIA) and clinical questionnaires. Results: A high expression of miR-5682 was associated with significantly lower values of BMI, fat mass, fat-free mass and plasma albumin at selected periods of RT course. miR-5682 allowed us to distinguish between patients classified with both SGA-C and low albumin level from other LC patients with 100% sensitivity and 69.6% specificity (AUC = 0.820; p < 0.0001). Higher expression of studied miRNA was significantly associated with shorter median overall survival (OS) in LC patients (HR = 2.26; p = 0.008). Conclusions: analysis of miR-5682 expression demonstrates a potential clinical utility in selection of LC patients suffering from nutritional deficiencies developing as a consequence of RT-based therapy.
Full article
(This article belongs to the Special Issue Non-coding RNAs in Human Health and Disease)
Open AccessArticle
GhCLCc-1, a Chloride Channel Gene from Upland Cotton, Positively Regulates Salt Tolerance by Modulating the Accumulation of Chloride Ions
by
Wenhao Li, Siqi Gao, Yinghao Zhao, Yuchen Wu, Xiaona Li, Jianing Li, Wei Zhu, Zongbin Ma and Wei Liu
Genes 2024, 15(5), 555; https://doi.org/10.3390/genes15050555 (registering DOI) - 26 Apr 2024
Abstract
The ionic toxicity induced by salinization has adverse effects on the growth and development of crops. However, researches on ionic toxicity and salt tolerance in plants have focused primarily on cations such as sodium ions (Na+), with very limited studies on
[...] Read more.
The ionic toxicity induced by salinization has adverse effects on the growth and development of crops. However, researches on ionic toxicity and salt tolerance in plants have focused primarily on cations such as sodium ions (Na+), with very limited studies on chloride ions (Cl−). Here, we cloned the homologous genes of Arabidopsis thaliana AtCLCc, GhCLCc-1A/D, from upland cotton (Gossypium hirsutum), which were significantly induced by NaCl or KCl treatments. Subcellular localization showed that GhCLCc-1A/D were both localized to the tonoplast. Complementation of Arabidopsis atclcc mutant with GhCLCc-1 rescued its salt-sensitive phenotype. In addition, the silencing of the GhCLCc-1 gene led to an increased accumulation of Cl− in the roots, stems, and leaves of cotton seedlings under salt treatments, resulting in compromised salt tolerance. And ectopic expression of the GhCLCc-1 gene in Arabidopsis reduced the accumulation of Cl− in transgenic lines under salt treatments, thereby enhancing salt tolerance. These findings elucidate that GhCLCc-1 positively regulates salt tolerance by modulating Cl− accumulation and could be a potential target gene for improving salt tolerance in plants.
Full article
(This article belongs to the Special Issue Cotton Genes, Genetics, and Genomics)
Open AccessArticle
Exploring the Role of the MUTYH Gene in Breast, Ovarian and Endometrial Cancer
by
Carla Lintas, Benedetta Canalis, Alessia Azzarà, Giovanna Sabarese, Giuseppe Perrone and Fiorella Gurrieri
Genes 2024, 15(5), 554; https://doi.org/10.3390/genes15050554 - 26 Apr 2024
Abstract
Background: MUTYH germline monoallelic variants have been detected in a number of patients affected by breast/ovarian cancer or endometrial cancer, suggesting a potential susceptibility role, though their significance remains elusive since the disease mechanism is normally recessive. Hence, the aim of this research
[...] Read more.
Background: MUTYH germline monoallelic variants have been detected in a number of patients affected by breast/ovarian cancer or endometrial cancer, suggesting a potential susceptibility role, though their significance remains elusive since the disease mechanism is normally recessive. Hence, the aim of this research was to explore the hypothesis that a second hit could have arisen in the other allele in the tumor tissue. Methods: we used Sanger sequencing and immunohistochemistry to search for a second MUTYH variant in the tumoral DNA and to assess protein expression, respectively. Results: we detected one variant of unknown significance, one variant with conflicting interpretation of pathogenicity and three benign/likely benign variants; the MUTYH protein was not detected in the tumor tissue of half of the patients, and in others, its expression was reduced. Conclusions: our results fail to demonstrate that germinal monoallelic MUTYH variants increase cancer risk through a LOH (loss of heterozygosity) mechanism in the somatic tissue; however, the absence or partial loss of the MUTYH protein in many tumors suggests its dysregulation regardless of MUTYH genetic status.
Full article
(This article belongs to the Special Issue Genetics of Multifactorial Diseases)
Open AccessReview
Exploring the Role of Cell-Free Nucleic Acids and Peritoneal Dialysis: A Narrative Review
by
Niccolò Morisi, Grazia Maria Virzì, Marco Ferrarini, Gaetano Alfano, Monica Zanella, Claudio Ronco and Gabriele Donati
Genes 2024, 15(5), 553; https://doi.org/10.3390/genes15050553 - 26 Apr 2024
Abstract
Introduction: Cell-free nucleic acids (cf-NAs) represent a promising biomarker of various pathological and physiological conditions. Since its discovery in 1948, cf-NAs gained prognostic value in oncology, immunology, and other relevant fields. In peritoneal dialysis (PD), blood purification is performed by exposing the
[...] Read more.
Introduction: Cell-free nucleic acids (cf-NAs) represent a promising biomarker of various pathological and physiological conditions. Since its discovery in 1948, cf-NAs gained prognostic value in oncology, immunology, and other relevant fields. In peritoneal dialysis (PD), blood purification is performed by exposing the peritoneal membrane. Relevant sections: Complications of PD such as acute peritonitis and peritoneal membrane aging are often critical in PD patient management. In this review, we focused on bacterial DNA, cell-free DNA, mitochondrial DNA (mtDNA), microRNA (miRNA), and their potential uses as biomarkers for monitoring PD and its complications. For instance, the isolation of bacterial DNA in early acute peritonitis allows bacterial identification and subsequent therapy implementation. Cell-free DNA in peritoneal dialysis effluent (PDE) represents a marker of stress of the peritoneal membrane in both acute and chronic PD complications. Moreover, miRNA are promising hallmarks of peritoneal membrane remodeling and aging, even before its manifestation. In this scenario, with multiple cytokines involved, mtDNA could be considered equally meaningful to determine tissue inflammation. Conclusions: This review explores the relevance of cf-NAs in PD, demonstrating its promising role for both diagnosis and treatment. Further studies are necessary to implement the use of cf-NAs in PD clinical practice.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
Open AccessArticle
Novel Genome-Engineered H Alleles Differentially Affect Lateral Inhibition and Cell Dichotomy Processes during Bristle Organ Development
by
Tanja C. Mönch, Thomas K. Smylla, Franziska Brändle, Anette Preiss and Anja C. Nagel
Genes 2024, 15(5), 552; https://doi.org/10.3390/genes15050552 - 26 Apr 2024
Abstract
Hairless (H) encodes the major antagonist in the Notch signaling pathway, which governs cellular differentiation of various tissues in Drosophila. By binding to the Notch signal transducer Suppressor of Hairless (Su(H)), H assembles repressor complexes onto Notch target genes. Using genome engineering,
[...] Read more.
Hairless (H) encodes the major antagonist in the Notch signaling pathway, which governs cellular differentiation of various tissues in Drosophila. By binding to the Notch signal transducer Suppressor of Hairless (Su(H)), H assembles repressor complexes onto Notch target genes. Using genome engineering, three new H alleles, HFA, HLLAAand HWA were generated and a phenotypic series was established by several parameters, reflecting the residual H-Su(H) binding capacity. Occasionally, homozygous HWA flies develop to adulthood. They were compared with the likewise semi-viable HNN allele affecting H-Su(H) nuclear entry. The H homozygotes were short-lived, sterile and flightless, yet showed largely normal expression of several mitochondrial genes. Typical for H mutants, both HWA and HNN homozygous alleles displayed strong defects in wing venation and mechano-sensory bristle development. Strikingly, however, HWA displayed only a loss of bristles, whereas bristle organs of HNN flies showed a complete shaft-to-socket transformation. Apparently, the impact of HWA is restricted to lateral inhibition, whereas that of HNN also affects the respective cell type specification. Notably, reduction in Su(H) gene dosage only suppressed the HNN bristle phenotype, but amplified that of HWA. We interpret these differences as to the role of H regarding Su(H) stability and availability.
Full article
(This article belongs to the Special Issue Gene Editing in Drosophila to Study Gene Function and Developmental Processes)
Open AccessArticle
Whole-Genome Resequencing Revealed Selective Signatures for Growth Traits in Hu and Gangba Sheep
by
Peifu Yang, Mingyu Shang, Jingjing Bao, Tianyi Liu, Jinke Xiong, Jupeng Huang, Jinghua Sun and Li Zhang
Genes 2024, 15(5), 551; https://doi.org/10.3390/genes15050551 - 26 Apr 2024
Abstract
A genomic study was conducted to uncover the selection signatures in sheep that show extremely significant differences in growth traits under the same breed, age in months, nutrition level, and management practices. Hu sheep from Gansu Province and Gangba sheep from the Tibet
[...] Read more.
A genomic study was conducted to uncover the selection signatures in sheep that show extremely significant differences in growth traits under the same breed, age in months, nutrition level, and management practices. Hu sheep from Gansu Province and Gangba sheep from the Tibet Autonomous Region in China were selected. We collected whole-genome data from 40 sheep individuals (24 Hu sheep and 16 Gangba sheep), through whole-genome sequencing. Selection signals were analyzed using parameters such as FST, π ratio, and Tajima’s D. We have identified several candidate genes that have undergone strong selection, particularly those associated with growth traits. Specifically, five growth-related genes were identified in both the Hu sheep group (HDAC1, MYH7B, LCK, ACVR1, GNAI2) and the Gangba sheep group (RBBP8, ACSL3, FBXW11, PLAT, CRB1). Additionally, in a genomic region strongly selected in both the Hu and Gangba sheep groups (Chr 22: 51,425,001-51,500,000), the growth-associated gene CYP2E1 was identified, further highlighting the genetic factors influencing growth characteristics in these breeds. This study analyzes the genetic basis for significant differences in sheep phenotypes, identifies candidate genes related to sheep growth traits, lays the foundation for molecular genetic breeding in sheep, and accelerates the genetic improvement in livestock.
Full article
(This article belongs to the Section Animal Genetics and Genomics)
►▼
Show Figures
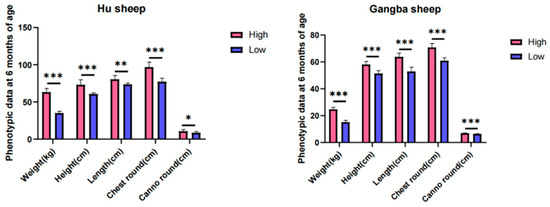
Figure 1
Open AccessArticle
Complete Chloroplast Genome Sequence Structure and Phylogenetic Analysis of Kohlrabi (Brassica oleracea var. gongylodes L.)
by
Mengliang Zhao, Yanxun Wu and Yanjing Ren
Genes 2024, 15(5), 550; https://doi.org/10.3390/genes15050550 - 26 Apr 2024
Abstract
Kohlrabi is an important swollen-stem cabbage variety belonging to the Brassicaceae family. However, few complete chloroplast genome sequences of this genus have been reported. Here, a complete chloroplast genome with a quadripartite cycle of 153,364 bp was obtained. A total of 132 genes
[...] Read more.
Kohlrabi is an important swollen-stem cabbage variety belonging to the Brassicaceae family. However, few complete chloroplast genome sequences of this genus have been reported. Here, a complete chloroplast genome with a quadripartite cycle of 153,364 bp was obtained. A total of 132 genes were identified, including 87 protein-coding genes, 37 transfer RNA genes and eight ribosomal RNA genes. The base composition analysis showed that the overall GC content was 36.36% of the complete chloroplast genome sequence. Relative synonymous codon usage frequency (RSCU) analysis showed that most codons with values greater than 1 ended with A or U, while most codons with values less than 1 ended with C or G. Thirty-five scattered repeats were identified and most of them were distributed in the large single-copy (LSC) region. A total of 290 simple sequence repeats (SSRs) were found and 188 of them were distributed in the LSC region. Phylogenetic relationship analysis showed that five Brassica oleracea subspecies were clustered into one group and the kohlrabi chloroplast genome was closely related to that of B. oleracea var. botrytis. Our results provide a basis for understanding chloroplast-dependent metabolic studies and provide new insight for understanding the polyploidization of Brassicaceae species.
Full article
(This article belongs to the Special Issue Plant Plastid Genome and Phylogenetics)
►▼
Show Figures
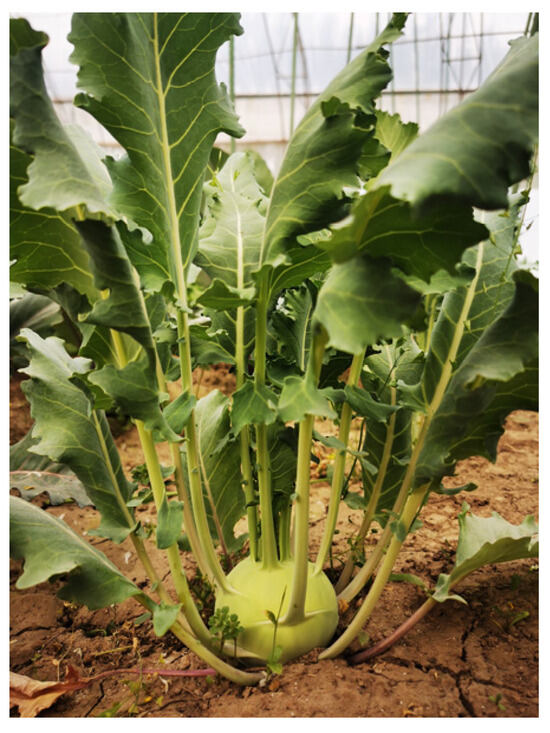

Figure 1
Open AccessArticle
A Bidirectional Non-Coding RNA Promoter Mediates Long-Range Gene Expression Regulation
by
Carlos Alberto Peralta-Alvarez, Hober Nelson Núñez-Martínez, Ángel Josué Cerecedo-Castillo, Augusto César Poot-Hernández, Gustavo Tapia-Urzúa, Sylvia Garza-Manero, Georgina Guerrero and Félix Recillas-Targa
Genes 2024, 15(5), 549; https://doi.org/10.3390/genes15050549 - 25 Apr 2024
Abstract
Recent evidence suggests that human gene promoters display gene expression regulatory mechanisms beyond the typical single gene local transcription modulation. In mammalian genomes, genes with an associated bidirectional promoter are abundant; bidirectional promoter architecture serves as a regulatory hub for a gene pair
[...] Read more.
Recent evidence suggests that human gene promoters display gene expression regulatory mechanisms beyond the typical single gene local transcription modulation. In mammalian genomes, genes with an associated bidirectional promoter are abundant; bidirectional promoter architecture serves as a regulatory hub for a gene pair expression. However, it has been suggested that its contribution to transcriptional regulation might exceed local transcription initiation modulation. Despite their abundance, the functional consequences of bidirectional promoter architecture remain largely unexplored. This work studies the long-range gene expression regulatory role of a long non-coding RNA gene promoter using chromosome conformation capture methods. We found that this particular bidirectional promoter contributes to distal gene expression regulation in a target-specific manner by establishing promoter–promoter interactions. In particular, we validated that the promoter–promoter interactions of this regulatory element with the promoter of distal gene BBX contribute to modulating the transcription rate of this gene; removing the bidirectional promoter from its genomic context leads to a rearrangement of BBX promoter–enhancer interactions and to increased gene expression. Moreover, long-range regulatory functionality is not directly dependent on its associated non-coding gene pair expression levels.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
Open AccessArticle
Non-Specific Epileptic Activity, EEG, and Brain Imaging in Loss of Function Variants in SATB1: A New Case Report and Review of the Literature
by
Flavia Privitera, Stefano Pagano, Camilla Meossi, Roberta Battini, Emanuele Bartolini, Domenico Montanaro and Filippo Maria Santorelli
Genes 2024, 15(5), 548; https://doi.org/10.3390/genes15050548 - 25 Apr 2024
Abstract
SATB1 (MIM #602075) is a relatively new gene reported only in recent years in association with neurodevelopmental disorders characterized by variable facial dysmorphisms, global developmental delay, poor or absent speech, altered electroencephalogram (EEG), and brain abnormalities on imaging. To date about thirty variants
[...] Read more.
SATB1 (MIM #602075) is a relatively new gene reported only in recent years in association with neurodevelopmental disorders characterized by variable facial dysmorphisms, global developmental delay, poor or absent speech, altered electroencephalogram (EEG), and brain abnormalities on imaging. To date about thirty variants in forty-four patients/children have been described, with a heterogeneous spectrum of clinical manifestations. In the present study, we describe a new patient affected by mild intellectual disability, speech disorder, and non-specific abnormalities on EEG and neuroimaging. Family studies identified a new de novo frameshift variant c.1818delG (p.(Gln606Hisfs*101)) in SATB1. To better define genotype–phenotype associations in the different types of reported SATB1 variants, we reviewed clinical data from our patient and from the literature and compared manifestations (epileptic activity, EEG abnormalities and abnormal brain imaging) due to missense variants versus those attributable to loss-of-function/premature termination variants. Our analyses showed that the latter variants are associated with less severe, non-specific clinical features when compared with the more severe phenotypes due to missense variants. These findings provide new insights into SATB1-related disorders.
Full article
(This article belongs to the Section Genetic Diagnosis)
Open AccessArticle
Statistical Genetic Approaches to Investigate Genotype-by-Environment Interaction: Review and Novel Extension of Models
by
Vincent P. Diego, Eron G. Manusov, Marcio Almeida, Sandra Laston, David Ortiz, John Blangero and Sarah Williams-Blangero
Genes 2024, 15(5), 547; https://doi.org/10.3390/genes15050547 - 25 Apr 2024
Abstract
Statistical genetic models of genotype-by-environment (G×E) interaction can be divided into two general classes, one on G×E interaction in response to dichotomous environments (e.g., sex, disease-affection status, or presence/absence of an exposure) and the other in response to continuous environments (e.g., physical activity,
[...] Read more.
Statistical genetic models of genotype-by-environment (G×E) interaction can be divided into two general classes, one on G×E interaction in response to dichotomous environments (e.g., sex, disease-affection status, or presence/absence of an exposure) and the other in response to continuous environments (e.g., physical activity, nutritional measurements, or continuous socioeconomic measures). Here we develop a novel model to jointly account for dichotomous and continuous environments. We develop the model in terms of a joint genotype-by-sex (for the dichotomous environment) and genotype-by-social determinants of health (SDoH; for the continuous environment). Using this model, we show how a depression variable, as measured by the Beck Depression Inventory-II survey instrument, is not only underlain by genetic effects (as has been reported elsewhere) but is also significantly determined by joint G×Sex and G×SDoH interaction effects. This model has numerous applications leading to potentially transformative research on the genetic and environmental determinants underlying complex diseases.
Full article
(This article belongs to the Special Issue Statistical Genetics of Human Complex Traits)
►▼
Show Figures
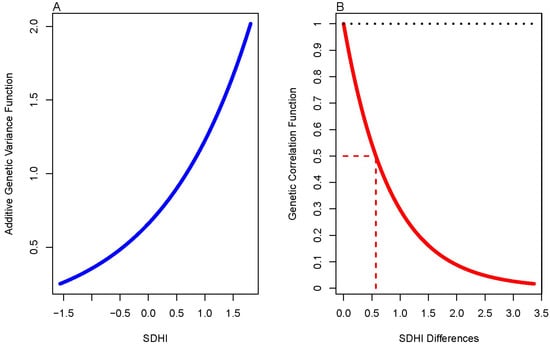
Figure 1

Journal Menu
► ▼ Journal Menu-
- Genes Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biology, BioMed, Cells, Genes
Applications of the Zebrafish Model
Topic Editors: De-Li Shi, Pengfei XuDeadline: 30 June 2024
Topic in
Biomedicines, Cells, CIMB, Genes, IJMS
Animal Models of Human Disease 2.0
Topic Editors: Sigrun Lange, Jameel M. InalDeadline: 31 August 2024
Topic in
Agriculture, Bioengineering, Genes, IJMS, Plants
Genetic Engineering in Agriculture
Topic Editors: Amy L. Klocko, Jianjun Chen, Haiwei LuDeadline: 30 September 2024
Topic in
Biology, BioMedInformatics, Cancers, Genes, IJMS
The 22nd International Conference on Bioinformatics (InCoB 2023): Translational Bioinformatics Transforming Life
Topic Editors: Jyotsna Batra, Srilakshmi Srinivasan, Shoba Ranganathan, Asif M. Khan, Harpreet SinghDeadline: 15 November 2024

Conferences
Special Issues
Special Issue in
Genes
Cotton Genes, Genetics, and Genomics
Guest Editor: Guanjing HuDeadline: 10 May 2024
Special Issue in
Genes
DNA Damage and Repair in Microorganisms, Plants and Mammalian Systems
Guest Editors: Ioly Kotta-Loizou, Nan Zhang, Xin WangDeadline: 20 May 2024
Special Issue in
Genes
Commemorating the Launch of the Section "Cytogenomics"
Guest Editor: Darren GriffinDeadline: 5 June 2024
Special Issue in
Genes
Genetic Markers and Liquid Biopsy for Kidney Diseases
Guest Editors: Guorong Li, Christophe MariatDeadline: 15 June 2024
Topical Collections
Topical Collection in
Genes
Study on Genotypes and Phenotypes of Pediatric Clinical Rare Diseases
Collection Editors: Livia Garavelli, Stefano Giuseppe Caraffi
Topical Collection in
Genes
Eukaryotic Non-coding RNAs: Diversity, Structure/Function, Implication in Cardiovascular Disease
Collection Editors: Morten Andre Høydal, Christiane Branlant
Topical Collection in
Genes
Feature Papers in ‘Animal Genetics and Genomics’
Collection Editors: Antonio Figueras, Raquel Vasconcelos








